anvi-profile-blitz
Table of Contents
FAST profiling of BAM files to get gene-, contig-, or genome-level coverage and detection stats. Unlike anvi-profile, which is another anvi'o program that can profile BAM files, this program is designed to be very quick and only report long-format files for various read recruitment statistics per item. Plase also see the program anvi-script-get-coverage-from-bam for recovery of data from BAM files without an anvi'o contigs database.
🔙 To the main page of anvi’o programs and artifacts.
Authors
Requires
Can use
collection-txt ![]() genes-of-interest-txt
genes-of-interest-txt ![]()
Provides
Usage
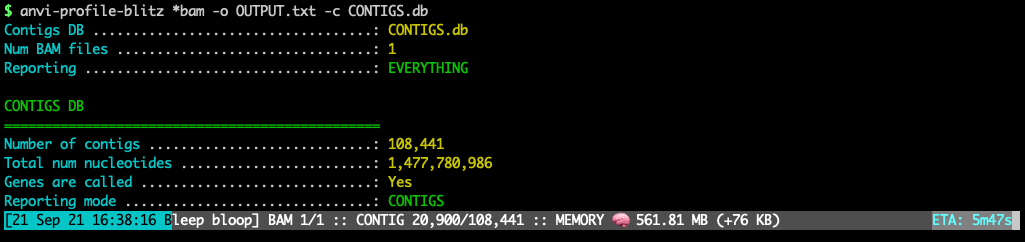
This program produces a bam-stats-txt from one or more bam-file given a contigs-db. It is designed to serve people who only need to process read recruitment data stored in a bam-file to recover coverage and detection statistics as well as the number of mapped reads (along with other statistics) for their genes, contigs, and/or genomes. It will report what’s going on nicely with memory usage information and estimated time of completion:
There are other programs in anvi’o software ecosystem that are similar to this one:
-
anvi-profile also takes a bam-file and profiles it. They both require a contigs-db. But while anvi-profile produces a single-profile-db for downstream analyses in anvi’o, anvi-profile-blitz produces text files for downstream analyses by the user (via R, Python, or other solutions). In contrast to anvi-profile, anvi-profile-blitz is orders of magnitude faster with similar memory usage.
-
anvi-script-get-coverage-from-bam also takes a bam-file and profiles it. They both produce text output files. But while anvi-script-get-coverage-from-bam does not require a contigs-db, anvi-profile-blitz requires one to work. They will both run very rapidly, anvi-script-get-coverage-from-bam will work with a much smaller amount of memory.
Output files
For output file formats, please see bam-stats-txt.
Running
You can use this program with one or more BAM files to recover minimal or extended statistics for contigs or genes in a contigs-db.
Since the program will not be able to ensure the contigs-db was generated from the same contigs-fasta that was used for read recruitment that resulted in bam-files for analysis, you can make serious mistakes if you mix up your workflow and start profiling BAM files that have nothing to do with a contigs-db. If you make a mistake like that, in the best case scenario you will get an empty output file because the program will skip all contigs with non-matching name. In the worst case scenario you will get a file if some names in contigs-db incorrectly matches to some names in the bam-file. While this warning may be confusing, you can avoid all these if you use the SAME FASTA FILE both as reference for read recruitment and as input for anvi-gen-contigs-database.
Contigs mode, default output
Profile contigs, produce a default output:
anvi-profile-blitz bam-file \ -c contigs-db \ -o OUTPUT.txt
This example is with a single BAM file, but you can also have multiple BAM files as a parameter by using wildcards,
anvi-profile-blitz *.bam \ -c contigs-db \ -o OUTPUT.txt
or by providing multiple paths:
anvi-profile-blitz /path/to/SAMPLE-01.bam \ /path/to/SAMPLE-02.bam \ /another/path/to/SAMPLE-03.bam -c contigs-db \ -o OUTPUT.txt
Contigs mode, minimal output
Profile contigs, produce a minimal output. This is the fastest option:
anvi-profile-blitz bam-file \ -c contigs-db \ --report-minimal \ -o OUTPUT.txt
Genome/bin mode via collections
If you have a collection-txt, you can summarize coverage/detection statistics per bin/genome instead of per contig.
anvi-profile-blitz bam-file \ -c contigs-db \ -C collection-txt \ -o OUTPUT.txt
Bin-level detection and coverage are computed by treating all contigs in a bin as a single genome (nucleotide arrays are concatenated under the hood). With --report-minimal, these statistics are streamed to keep memory usage close to contig mode even for large bins.
Genes mode, default output
Instead of contigs, profile genes, produce a default output:
anvi-profile-blitz bam-file \ -c contigs-db \ --gene-mode \ -o OUTPUT.txt
--report-minimal will behave the same, and produce minimal output.
Genes mode, on a subset of genes
You don’t want to profile all genes in your contigs-db? Then you should tell anvi’o which specific genes you are interested in by either providing a comma-separated list of gene-caller IDs:
the following will profile 4 specific gene calls
anvi-profile-blitz bam-file \ -c contigs-db \ --gene-mode \ -o OUTPUT.txt \ --gene-caller-ids 5,13,74,203
Or by providing a genes-of-interest-txt file:
anvi-profile-blitz bam-file \ -c contigs-db \ --gene-mode \ -o OUTPUT.txt \ --genes-of-interest genes-of-interest-txt
Performance
The memory use will be correlated linaerly with the size of the contigs-db, but once everything is loaded, the memory usage will not increase substantially over time.
With the flag --report-minimal, anvi-profile-blitz profiled on a laptop computer 100,000 contigs that contained 1 billion nts in 6 minutes and used ~300 Mb memory. This contigs database had 1.5 million genes, and memory usage increased to 1.7 Gb when anvi-profile-blitz run in --gene-mode. The flag --gene-mode does not change time complexity dramatically.
Anvi’o has this program because Emile Faure presented us with a challenge: Emile had a ~140 Gb anvi’o contigs-db that contained nearly 70 million contig sequences from over 200 single-assembled metagenomes, and wanted to learn the coverages of each gene in the contigs database in 200 metagenomes individually. Yet the combination of anvi-profile and anvi-summarize jobs would take more than 40 days to complete. Since all Emile needed was to learn the coverages from BAM files, we implemented anvi-profile-blitz to skip the profiling step. The run took 8 hours to compute and report coverage values for 175 million genes in 70 million contigs, and the memory use remained below 200 Gb.
Edit this file to update this information.
Additional Resources
Are you aware of resources that may help users better understand the utility of this program? Please feel free to edit this file on GitHub. If you are not sure how to do that, find the __resources__ tag in this file to see an example.