An anvi'o tutorial with Trichodesmium genomes
Table of Contents
Summary
The purpose of this tutorial is to learn how to use the set of integrated ‘omics tools in anvi’o to make sense of a few Trichodesmium genomes. Here is a list of topics that are covered in this tutorial:
- Create contigs databases and run functional annotation programs.
- Estimate taxonomy and completion/redundancy across multiple genomes.
- Generate a pangenome of closely related Trichodesmium genomes.
- Study metabolism by predicting metabolic pathway completeness and metabolic interactions.
Because it covers so many different topics, we’ve split up the tutorial into different chapters. You’ll find links to each chapter below.
If you have any questions about this exercise, or have ideas to make it better, please feel free to get in touch with the anvi’o community through our Discord server:
To reproduce these exercises with your own dataset, you should first follow the instructions here to install anvi’o.
A story of nitrogen fixation (or not) in Trichodesmium
This tutorial will largely recapitulate a story from the following paper, published by Tom Delmont in 2021:
- Expanded the understanding of the ecological roles and diversity of Trichodesmium.
- Challenged long-held assumptions about nitrogen fixation in marine cyanobacteria.
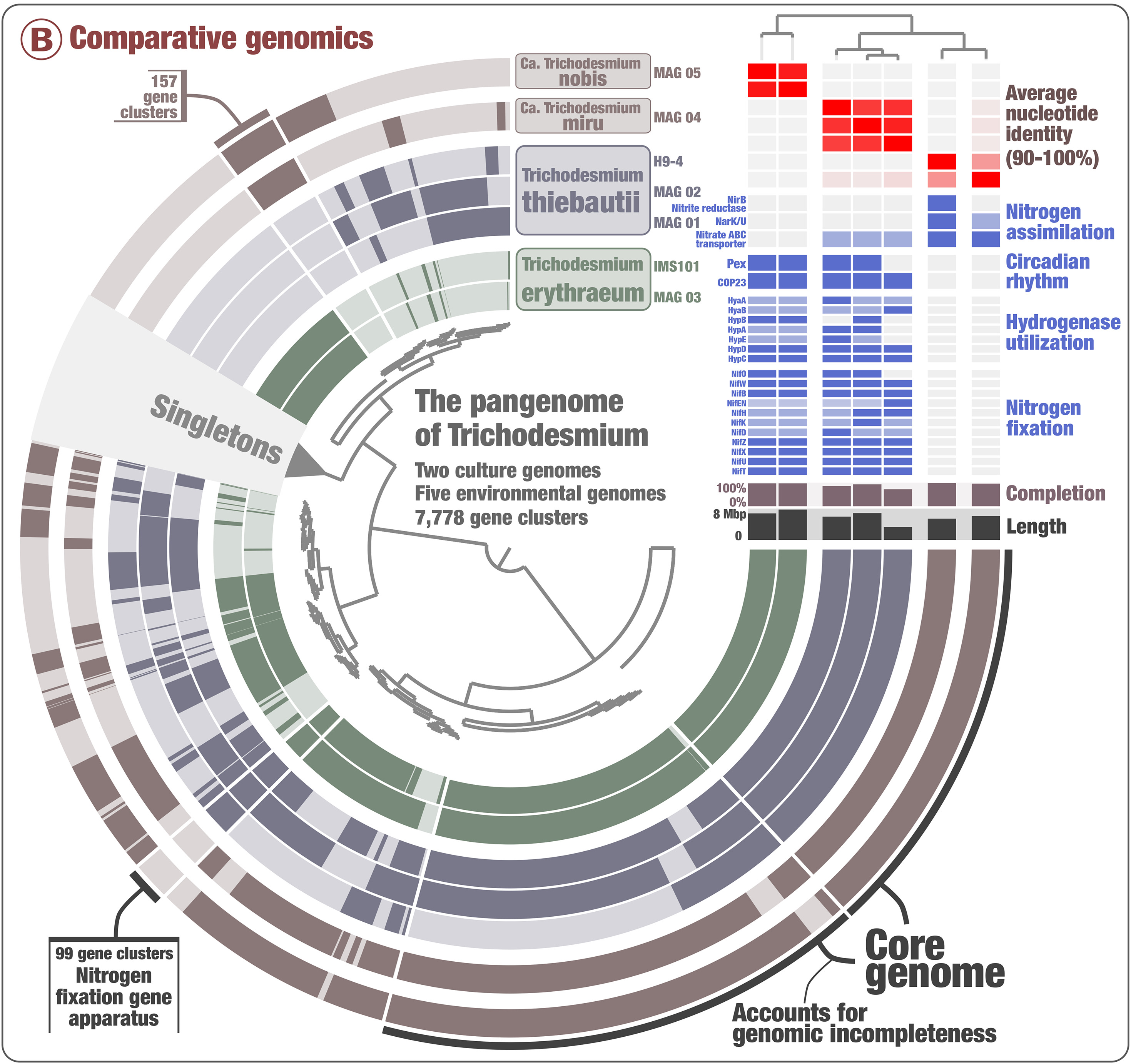
As a little preview, the essence of the story is this: Trichodesmium species are well-known cyanobacterial nitrogen fixers (‘diazotrophs’) in the global oceans, but – surprise – it turns out that not all of them can do nitrogen fixation. Tom used a combination of pangenomics, phylogenomics, and clever read recruitment analyses on a set of MAGs and reference genomes to demonstrate that two new (candidate) Trichodesmium species, Candidatus T. miru and Candidatus T. nobis, are nondiazotrophic.
We will use a variety of anvi’o programs to investigate the same genomes and characterize their nitrogen-fixing capabilities, to demonstrate how you, too, could discover cool microbial ecology stories like this one.
Downloading the datapack
In your terminal, choose a working directory for this tutorial and use the following code to download the dataset:
curl -L https://cloud.uol.de/public.php/dav/files/S67286XGxtax2AX \
-o trichodesmium_tutorial.tar.gz
Then unpack it, and go into the datapack directory:
tar -zxvf trichodesmium_tutorial.tar.gz
cd trichodesmium_tutorial
At this point, if you check the datapack contents in your terminal with ls, this is what you should be seeing:
$ ls
00_DATA
$ ls 00_DATA/
associate_dbs fasta metabolism_state.json module_info.txt nitrogen_heatmap.json pan_state.json
contigs genome-pairs.txt metagenome modules nitrogen_step_copies.json phylo_dbs
Inside the 00_DATA folder, there are several files that will be useful for various parts of this tutorial. We will start from the seven Trichodesmium genomes stored in the fasta directory. Some are metagenome-assembled genomes (MAGs) binned from the TARA Ocean metagenomic dataset, and others are reference genomes taken from NCBI RefSeq.
Activating anvi’o
Before you start, don’t forget to activate your anvi’o environment:
We use the development version of anvi’o here, but you could also use a stable release of anvi’o if that is what you have installed. Any stable release starting from v9 or later will include all of the programs covered in this tutorial. If you try an earlier release, you may see “command not found” errors for some of the commands.
conda activate anvio-dev
Tutorial Chapter Navigation
Each chapter of this tutorial has its own webpage and is technically independent. You can click on the links below to access the individual chapters. As long as you have downloaded the datapack from above, you’ll be able to work through any chapter regardless of whether you completed previous chapters.
- Tutorial introduction (main page) ← you are here
- Chapter 1: Genomics
- Chapter 2: Pangenomics
- Chapter 3: Phylogenomics
- Chapter 4: Metabolism
For convenience, you’ll find this set of links at the top of each chapter’s webpage.
This tutorial is still a work-in-progress. More sections coming soon!