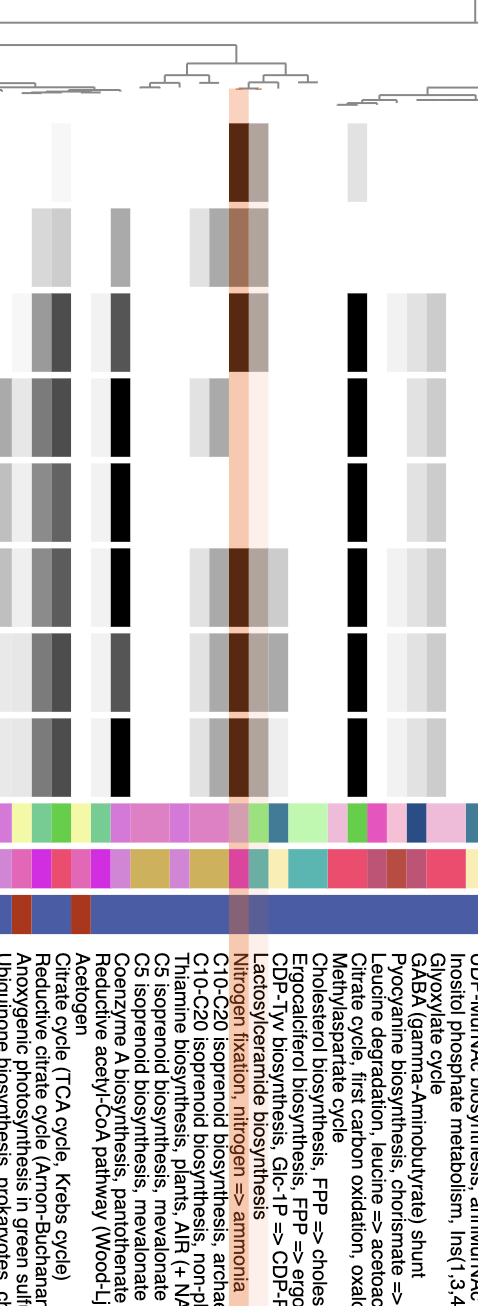An anvi'o tutorial with Trichodesmium genomes (Chapter 4)
Table of Contents
About this page This webpage is one chapter of a much larger effort to cover multiple aspects of anvi’o in the same tutorial. If you need more context, please visit the main page of the tutorial, where you will find information about the dataset we are working with and the commands to download the tutorial datapack.
Quick Navigation
- Tutorial introduction (main page)
- Chapter 1: Genomics
- Chapter 2: Pangenomics
- Chapter 3: Phylogenomics
- Chapter 4: Metabolism ← you are here
Metabolism
Show/Hide Starting the tutorial at this section? Click here for data preparation steps.
If you haven’t run previous sections of this tutorial (particularly the ‘Working with multiple genomes’ section), then you should follow these steps to setup the files that we will use going forward.
cp 00_DATA/contigs/*-contigs.db .
anvi-script-gen-genomes-file --input-dir . -o external-genomes.txt
ls 00_DATA/fasta | cut -d "." -f1 > genomes.txt
Looking at individual gene functions is undisputably useful for understanding the lifestyle and potential activities of microbes. However, most genes do not work in isolation. Metabolic capacities such as nitrogen fixation, photosynthesis, biosynthesis of cellular components, and carbon utilization require multiple enzymes (or enzyme components) working sequentially or in parallel to catalyze all chemical reactions in a so-called metabolic pathway. In many cases, there are also several possible versions of these enzymes – variants across different domains of life, taxonomic groups, or environmental conditions – that could be used to catalyze the same set of chemical reactions. Manually looking for all possible versions of all enzymes needed for a given metabolic capacity is possible (we just did it in the pangenomics section for nitrogen fixation), but it isn’t very efficient (and requires you to know exactly what you are looking for). And that sort of approach is unscaleable when you are interested in more than a handful of specific metabolic pathways.
This section of the tutorial covers metabolism reconstruction, a strategy for summarizing all the metabolic capacities of a given organism (or community) by examining functional annotations in their metabolic context. To do this, one can either aggregate the genes related to multiple individual metabolic pathways to evaluate the organism’s capacity to do specific, ecologically-relevant things (“pathway prediction”) or use all annotated enzymes to create the entire network of chemical reactions that the organism can catalyze (“metabolic modeling”).
To keep things organized, we’ll make a new subdirectory for ourselves in which we can work. We’ll copy the external genomes file in here for convenience.
mkdir -p 02_METABOLISM && cd 02_METABOLISM/
cp ../external-genomes.txt .
Estimating metabolic pathway completeness
We’ll start with pathway prediction. The program anvi-estimate-metabolism computes completeness scores (and copy numbers) of metabolic modules. By default, it uses modules from the KEGG MODULE database, which contains a lot of well-studied metabolic pathways of general interest that are defined in terms of KEGG Ortholog (KO) protein families. When you run anvi-setup-kegg-data to get the KEGG KOfam models used for annotation with anvi-run-kegg-kofams, you also set up the KEGG MODULE data on your computer.
Show/Hide What version of KEGG data do you have?
The KEGG database goes through regular updates, so to keep things a bit more stable anvi’o uses prepackaged snapshots of KEGG data. More reasons for this are explained here. You can have multiple different snapshots on your computer at the same time, and pick which one to use with the --kegg-data-dir parameter of KEGG-associated programs.
To keep track of which KEGG snapshot is relevant to your current dataset, anvi’o hashes the contents of the KEGG data directory and stores this hash in your contigs-db when you run anvi-run-kegg-kofams. You can see which version you have by running anvi-db-info and looking for the modules_db_hash key. For example, all of the Trichodesmium genomes in the tutorial datapack should have the following hash value:
modules_db_hash ..............................: 68221bd12b30
This hash enables us to ensure that the version of KEGG used to annotate your (meta)genome matches to the version used for metabolism reconstruction.
anvi-estimate-metabolism can work on individual genomes, but we’re interested in comparing the metabolic capacity of all 8 Trichodesmium genomes. So let’s use our handy-dandy external-genomes file to individually estimate metabolism on each one:
# takes ~30 seconds
anvi-estimate-metabolism -e external-genomes.txt -O tricho_metabolism
You should get an output file called tricho_metabolism_modules.txt that looks like this:
module |
genome_name |
db_name |
module_name |
module_class |
module_category |
module_subcategory |
module_definition |
stepwise_module_completeness |
stepwise_module_is_complete |
pathwise_module_completeness |
pathwise_module_is_complete |
proportion_unique_enzymes_present |
enzymes_unique_to_module |
unique_enzymes_hit_counts |
enzyme_hits_in_module |
gene_caller_ids_in_module |
warnings |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| M00001 | MAG_Candidatus_Trichodesmium_miru | MAG_Candidatus_Trichodesmium_miru | Glycolysis (Embden-Meyerhof pathway), glucose => pyruvate | Pathway modules | Carbohydrate metabolism | Central carbohydrate metabolism | “(K00844,K12407,K00845,K25026,K00886,K08074,K00918) (K01810,K06859,K13810,K15916) (K00850,K16370,K21071,K24182,K00918) (K01623,K01624,K11645,K16305,K16306) K01803 ((K00134,K00150) K00927,K11389) (K01834,K15633,K15634,K15635) (K01689,K27394) (K00873,K12406)” | 1.0 | True | 1.0 | True | NA | No enzymes unique to module | NA | K00134,K00134,K00845,K00845,K00873,K00927,K01623,K01624,K01689,K01803,K01803,K01810,K01834,K15633,K21071,K25026 | 2899,2579,4200,3691,985,2838,2319,1760,2136,4011,3621,2573,2274,2197,1300,1049 | K00134 is present in multiple modules: M00001/M00002/M00003/M00308/M00552/M00165/M00611/M00612,K00845 is present in multiple modules: M00001/M00549,K00873 is present in multiple modules: M00001/M00002,K00927 is present in multiple modules: M00001/M00002/M00003/M00308/M00552/M00165/M00611/M00612,K01623 is present in multiple modules: M00001/M00003/M00165/M00611/M00612,K01624 is present in multiple modules: M00001/M00003/M00165/M00345/M00344/M00611/M00612,K01689 is present in multiple modules: M00001/M00002/M00003/M00346,K01803 is present in multiple modules: M00001/M00002/M00003/M00165,K01810 is present in multiple modules: M00001/M00004,K01834 is present in multiple modules: M00001/M00002/M00003,K15633 is present in multiple modules: M00001/M00002/M00003,K21071 is present in multiple modules: M00001/M00345,K25026 is present in multiple modules: M00001/M00549 |
| M00002 | MAG_Candidatus_Trichodesmium_miru | MAG_Candidatus_Trichodesmium_miru | Glycolysis, core module involving three-carbon compounds | Pathway modules | Carbohydrate metabolism | Central carbohydrate metabolism | “K01803 ((K00134,K00150) K00927,K11389) (K01834,K15633,K15634,K15635) (K01689,K27394) (K00873,K12406)” | 1.0 | True | 1.0 | True | NA | No enzymes unique to module | NA | K00134,K00134,K00873,K00927,K01689,K01803,K01803,K01834,K15633 | 2899,2579,985,2838,2136,4011,3621,2274,2197 | K00134 is present in multiple modules: M00001/M00002/M00003/M00308/M00552/M00165/M00611/M00612,K00873 is present in multiple modules: M00001/M00002,K00927 is present in multiple modules: M00001/M00002/M00003/M00308/M00552/M00165/M00611/M00612,K01689 is present in multiple modules: M00001/M00002/M00003/M00346,K01803 is present in multiple modules: M00001/M00002/M00003/M00165,K01834 is present in multiple modules: M00001/M00002/M00003,K15633 is present in multiple modules: M00001/M00002/M00003 |
| M00003 | MAG_Candidatus_Trichodesmium_miru | MAG_Candidatus_Trichodesmium_miru | Gluconeogenesis, oxaloacetate => fructose-6P | Pathway modules | Carbohydrate metabolism | Central carbohydrate metabolism | “(K01596,K01610) (K01689,K27394) (K01834,K15633,K15634,K15635) K00927 (K00134,K00150) K01803 ((K01623,K01624,K11645) (K03841,K02446,K11532,K01086,K04041),K01622)” | 0.8571428571428571 | True | 0.875 | True | NA | No enzymes unique to module | NA | K00134,K00134,K00927,K01623,K01624,K01689,K01803,K01803,K01834,K03841,K15633 | 2899,2579,2838,2319,1760,2136,4011,3621,2274,18,2197 | K00134 is present in multiple modules: M00001/M00002/M00003/M00308/M00552/M00165/M00611/M00612,K00927 is present in multiple modules: M00001/M00002/M00003/M00308/M00552/M00165/M00611/M00612,K01623 is present in multiple modules: M00001/M00003/M00165/M00611/M00612,K01624 is present in multiple modules: M00001/M00003/M00165/M00345/M00344/M00611/M00612,K01689 is present in multiple modules: M00001/M00002/M00003/M00346,K01803 is present in multiple modules: M00001/M00002/M00003/M00165,K01834 is present in multiple modules: M00001/M00002/M00003,K03841 is present in multiple modules: M00003/M00165/M00344/M00611/M00612,K15633 is present in multiple modules: M00001/M00002/M00003 |
| M00307 | MAG_Candidatus_Trichodesmium_miru | MAG_Candidatus_Trichodesmium_miru | Pyruvate oxidation, pyruvate => acetyl-CoA | Pathway modules | Carbohydrate metabolism | Central carbohydrate metabolism | ”((K00163,K00161+K00162)+K00627+K00382-K13997),K00169+K00170+K00171+(K00172,K00189),K03737” | 1.0 | True | 1.0 | True | 1.0 | K00161,K00162,K00627 | 1,1,1 | K00161,K00162,K00382,K00627 | 2967,1381,2978,1001 | K00382 is present in multiple modules: M00307/M00009/M00011/M00532/M00621/M00036/M00032 |
| M00009 | MAG_Candidatus_Trichodesmium_miru | MAG_Candidatus_Trichodesmium_miru | Citrate cycle (TCA cycle, Krebs cycle) | Pathway modules | Carbohydrate metabolism | Central carbohydrate metabolism | “(K01647,K05942,K01659) (K01681,K27802,K01682) (K00031,K00030) ((K00164+K00658,K01616)+K00382,K00174+K00175-K00177-K00176) (K01902+K01903,K01899+K01900,K18118) (K00234+K00235+K00236+(K00237,K25801),K00239+K00240+K00241-(K00242,K18859,K18860),K00244+K00245+K00246-K00247) (K01676,K01679,K01677+K01678) (K00026,K00025,K00024,K00116)” | 0.625 | False | 0.7708333333333334 | True | NA | No enzymes unique to module | NA | K00031,K00239,K00240,K00382,K01659,K01679,K01682,K01902,K01903 | 3698,3854,3955,2978,1925,1837,3626,2385,2386 | K00031 is present in multiple modules: M00009/M00010/M00740/M00173/M00614,K00239 is present in multiple modules: M00009/M00011/M00982/M00173/M00376/M00374/M00149/M00613/M00614,K00240 is present in multiple modules: M00009/M00011/M00982/M00173/M00376/M00374/M00149/M00613/M00614,K00382 is present in multiple modules: M00307/M00009/M00011/M00532/M00621/M00036/M00032,K01659 is present in multiple modules: M00009/M00010/M00012/M00740/M00982,K01679 is present in multiple modules: M00009/M00011/M00982/M00173/M00376/M00613/M00614,K01682 is present in multiple modules: M00009/M00010/M00012/M00982/M00173/M00614,K01902 is present in multiple modules: M00009/M00011/M00173/M00374/M00620/M00614,K01903 is present in multiple modules: M00009/M00011/M00173/M00374/M00620/M00614 |
An explanation of the output columns can be found on the kegg-metabolism help page. There, you will also see the other possible output types you could request using the --output-modes flag.
To stick with the nitrogen fixation theme we’ve been following so far, let’s look for nitrogen fixation in this file. KEGG’s nitrogen fixation module is M00175. To have this module complete, a genome has to include either the nifHDK genes of the molybdenum-dependent nitrogenase enzyme complex, OR the vnfDKGH genes of the vanadium-dependent nitrogenase complex. This module unfortunately does not include other necessary nitrogen fixation genes (as Iva has complained about before), but it is a good enough start.
There are many ways to search for the nitrogen fixation module and look at its completeness scores within each genome. Here is one way using BASH:
head -n 1 tricho_metabolism_modules.txt | cut -f 1,2,9,11 > nif_table.txt
grep -i "nitrogen fixation" tricho_metabolism_modules.txt | cut -f 1,2,9,11 >> nif_table.txt
And here is what the resulting table looks like:
module |
genome_name |
stepwise_module_completeness |
pathwise_module_completeness |
|---|---|---|---|
| M00175 | MAG_Trichodesmium_erythraeum | 1.0 | 1.0 |
| M00175 | MAG_Trichodesmium_thiebautii_Atlantic | 0.0 | 0.6666666666666666 |
| M00175 | MAG_Trichodesmium_thiebautii_Indian | 1.0 | 1.0 |
| M00175 | Trichodesmium_erythraeum_IMS101 | 1.0 | 1.0 |
| M00175 | Trichodesmium_sp | 1.0 | 1.0 |
| M00175 | Trichodesmium_thiebautii_H9_4 | 1.0 | 1.0 |
The first thing to notice is that T. miru and T. nobis are not in the table at all. This implies that their completeness scores for this module were both 0.0, since by default anvi-estimate-metabolism doesn’t include these zero results in the output to save on space (if you want those zero values to be in the table, you could use the flag --include-zeros). Okay, so that matches up to our expectations so far.
The second thing to notice is that there are two types of completeness score, stepwise and pathwise. A full explanation of these metrics can be found here. The short version is that pathwise completeness considers all possible combinations of enzymes that fullfill the module (in this case, either nifHDK or vnfDKGH) and computes the percentage of enzymes annotated for each combination (reporting the maximum), while stepwise completeness breaks down the module into overall steps and only considers each step complete if all enzymes required for the step are present. Nitrogen fixation is just one reaction requiring an enzyme complex made up of multiple parts, so the stepwise interpretation of the module considers it to be just one step and if not all of the enzyme components are present, that step is incomplete. This is why the stepwise completeness for MAG_Trichodesmium_thiebautii_Atlantic is 0.0 while the pathwise completeness is 0.66 – MAG_Trichodesmium_thiebautii_Atlantic is missing one of the required enzyme components. In fact, if you look at the enzyme_hits_in_module column for this MAG and this module, you will see that only K02588 (nifH) and K02591 (nifK) are annotated. K02586 (nifD) is missing.
Comparing metabolic capacity across genomes
It’s great to have all the details about each individual metabolic module in each genome. But it is a lot of information to parse through, and sometimes we just want a nice picture to look at. So let’s repeat the metabolism estimation analysis, but this time let’s ask for matrix-type output that we can easily visualize in the anvi’o interactive interface:
# takes ~10 seconds
anvi-estimate-metabolism -e external-genomes.txt -O tricho_metabolism --matrix-format
You should get a bunch of different output files, but the one we will visualize is the matrix of pathwise completeness scores: tricho_metabolism-module_pathwise_completeness-MATRIX.txt. We can give this file to the interactive interface in --manual mode, along with the name of a (to be created) profile-db to store the interface settings:
anvi-interactive -d tricho_metabolism-module_pathwise_completeness-MATRIX.txt \
-p metabolism_profile.db \
--manual \
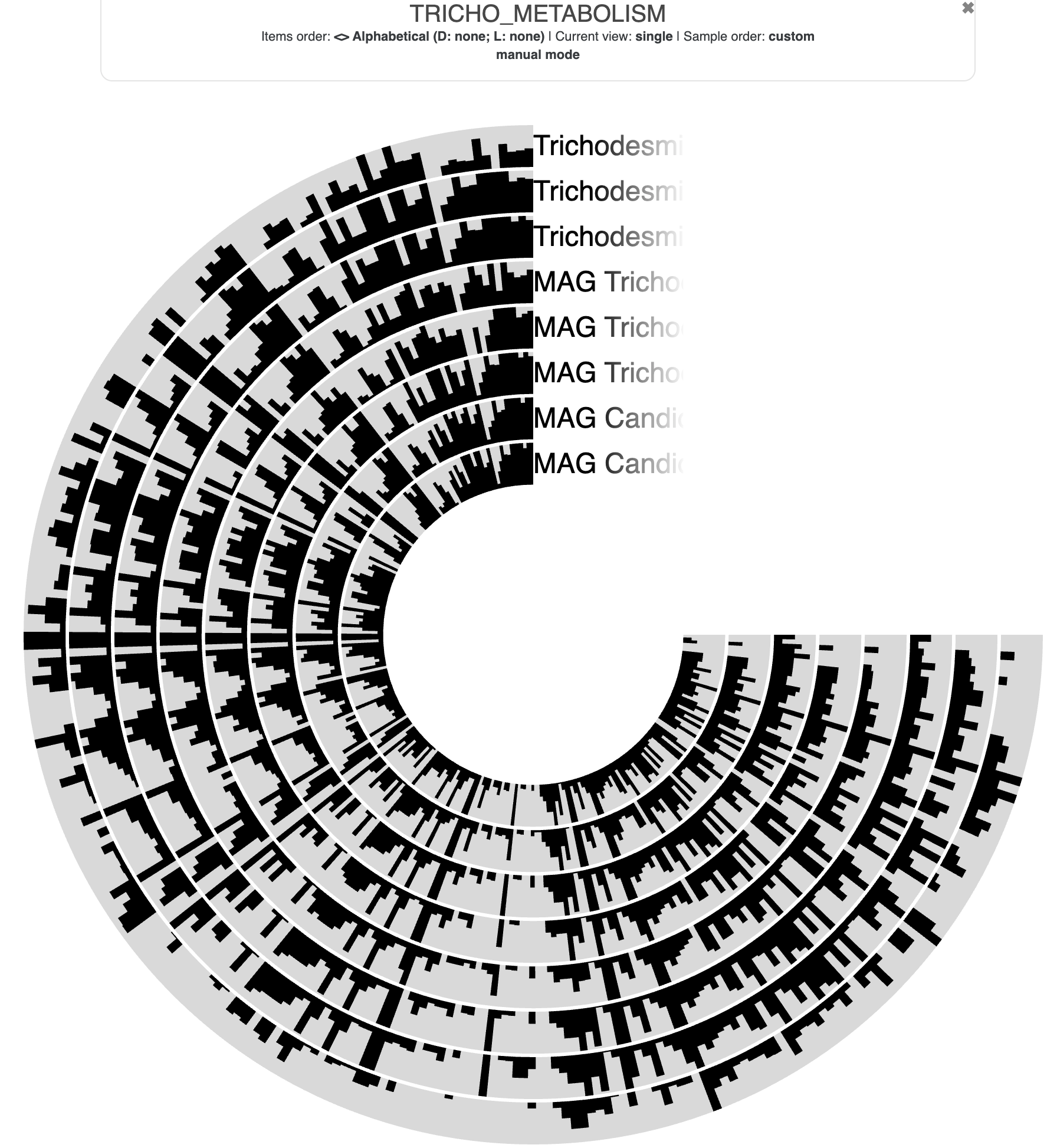
--title TRICHO_METABOLISM
The resulting visualization will look something like this:
We can make it much easier to see differences between the genomes by doing the following things:
- making the visualization rectangular (‘Drawing Type’ => ‘Phylogram’) and bigger (in ‘Options’ tab, increase Dendrogram Width)
- changing the bar plots into a heatmap (select ‘Intensity’ chart type in the Items Display settings)
- playing with the ‘Min’ values to filter out highly incomplete modules (also under Items Display)
- clustering the genomes so that genomes with similar metabolic capacity are close together
- clustering the modules so that modules with similar distribution across genomes are close together
- importing the name and categories of each module
The first three things can be done by tweaking the interface settings. And if you click on the dropdown box below, you can see all the terminal steps for clustering and importing.
Show/Hide Organizing the metabolism heatmap
Here is how to cluster the modules (rows of the matrix, which become ‘items’ in the interface):
anvi-matrix-to-newick tricho_metabolism-module_pathwise_completeness-MATRIX.txt
This tree file can be given directly to anvi-interactive using the -t parameter. Or, you could import it into the profile-db as an ‘items order’ using the program anvi-import-items-order. We’ll stick to the former in this tutorial.
Here is how to cluster the genomes (columns of the matrix, which become ‘layers’ in the interface). First, you flip the matrix to put the genomes into the rows, and then you use the same anvi-matrix-to-newick command as before:
anvi-script-transpose-matrix tricho_metabolism-module_pathwise_completeness-MATRIX.txt -o tricho_metabolism-module_pathwise_completeness-MATRIX-transposed.txt
anvi-matrix-to-newick tricho_metabolism-module_pathwise_completeness-MATRIX-transposed.txt
Then, you put the resulting dendrogram into a misc-data-layer-orders-txt file:
# read the file into a variable, and then print to a new tab-delimited file
TREE=$(cat tricho_metabolism-module_pathwise_completeness-MATRIX-transposed.txt.newick)
echo -e "item_name\tdata_type\tdata_value\nmag_organization\tnewick\t$TREE" > layer_order.txt
This allows you to import the dendrogram into the profile-db:
anvi-import-misc-data -p metabolism_profile.db -t layer_orders \
layer_order.txt
Finally, we want to see module information like names and categories, not just the module numbers. Here’s a little set of SQL queries to extract that information from the modules-db (use the database from the same KEGG data directory you’ve been using all along):
# if you aren't using the default KEGG data dir, you should change this variable to point to the MODULES.db in the dir you ARE using
export ANVIO_MODULES_DB=`python -c "import anvio; import os; print(os.path.join(os.path.dirname(anvio.__file__), 'data/misc/KEGG/MODULES.db'))"`
sqlite3 $ANVIO_MODULES_DB "select module,data_value from modules where data_name='NAME'" | \
tr '|' '\t' > module_names.txt
sqlite3 $ANVIO_MODULES_DB "select module,data_value from modules where data_name='CLASS'" | \
tr '|' '\t' > module_categories.txt
You can split the category strings into 3 different columns, and combine everything into one table:
echo -e "class\tcategory\tsubcategory" > category_columns.txt
cut -f 2 module_categories.txt | sed 's/; /\t/g' >> category_columns.txt
echo -e "module\tname" > name_columns.txt
cat module_names.txt >> name_columns.txt
paste name_columns.txt category_columns.txt > module_info.txt
Then you can import that new table into the profile-db:
anvi-import-misc-data -t items -p metabolism_profile.db module_info.txt
## clean up
rm module_names.txt module_categories.txt name_columns.txt category_columns.txt
Now you should have everything you need for visualizing the data nicely.
Once you are finished with the code in the dropdown box, you can visualize the pathwise completeness matrix again like this (adding the module organization with the -t parameter):
anvi-interactive -d tricho_metabolism-module_pathwise_completeness-MATRIX.txt \
-p metabolism_profile.db \
--manual \
--title TRICHO_METABOLISM \
-t tricho_metabolism-module_pathwise_completeness-MATRIX.txt.newick
In case you want your visualization to exactly match ours, you can import our settings into the profile-db for the heatmap. Note that the organization will only work if you named your trees the same way we did.
anvi-import-state -s ../00_DATA/metabolism_state.json \
-p metabolism_profile.db \
-n default
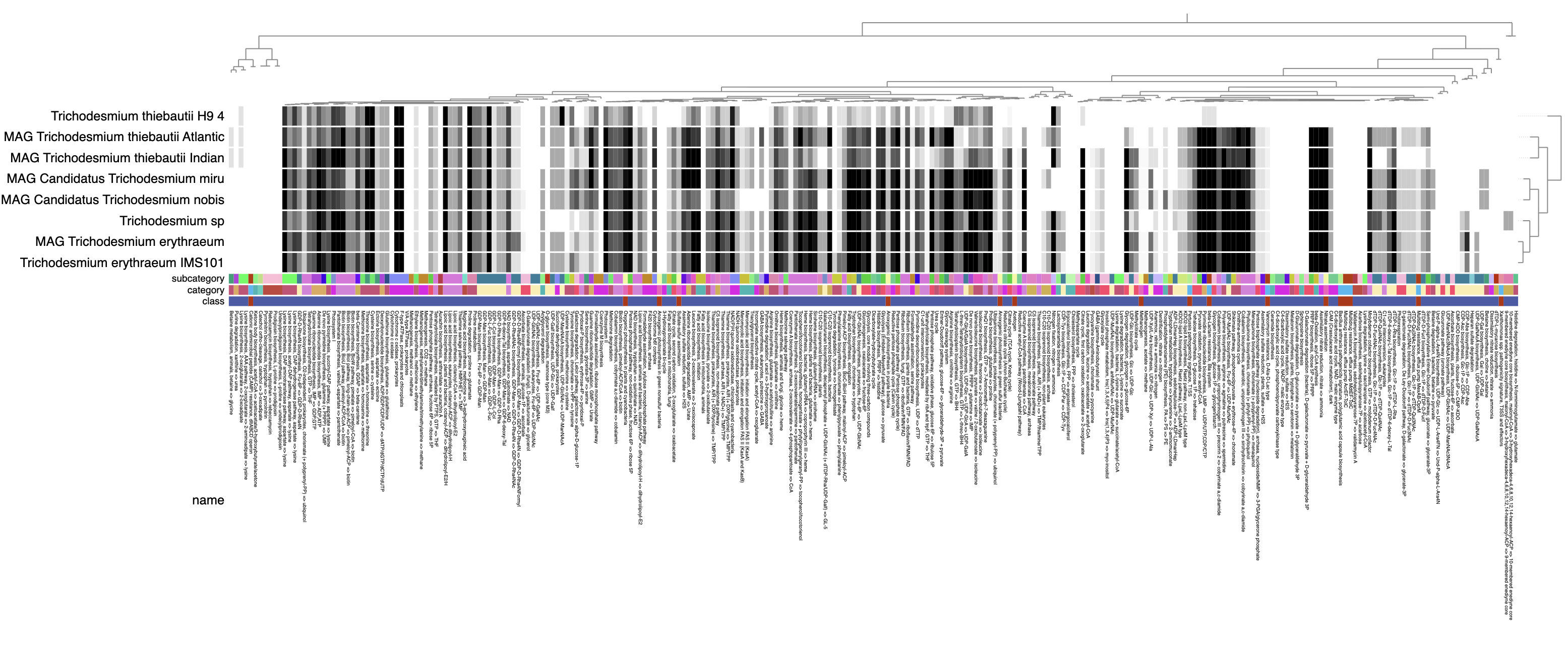
The heatmap should ultimately look something like this:
A few things we can notice from the visualization:
- In the genome clustering, there are three main groups of metabolically-similar genomes: all the T. erythraeum genomes are together, all the T. thiebautii genomes are together, and the two candidate species T. miru and T. nobis are together.
- The rather incomplete T. thiebautii H9 genome is clearly missing a lot of metabolic pathways that it shouldn’t actually be missing, particularly on the right side of the heatmap where there are plenty of modules shared by all of the other 7 genomes.
- It should be fairly easy now to spot the modules that are missing specifically in T. miru and T. nobis. Here they are, highlighted in orange:
As expected, the nitrogen fixation module is there.
Right next to the nitrogen fixation module is an unusual sounding metabolic pathway, Lactosylceramide biosynthesis, which is 50% complete in all of the other genomes. Lactosylceramides are a type of glycosphingolipid, not very well studied in Cyanobacteria (except for some work investigating sphingolipid roles in plant-microbe symbiotic interactions (Heaver, Johnson and Ley 2018)), so perhaps it is not this exact pathway that is relevant, but rather some of the enzymes in it. Indeed, if you look at the details of M00066 in the long-format output file, you will see that only one enzyme is annotated in these genomes: ceramide glucosyltransferase (K00720). Unfortunately, this enzyme is also not well studied in bacteria, so we don’t have much literature backup for interpreting the lack of this function in T. miru and T. nobis. Maybe a sphingolipid expert will see this one day and look into it. :)
Using custom metabolic modules
When we estimated metabolism with the default KEGG modules in the previous section, a lot of the completeness scores were quite similar across all 8 of our genomes. But we know from Tom’s paper that there are a few other nitrogen fixation-related activities that T. miru and T. nobis do not have and that the other genomes do. The KEGG modules don’t really capture what makes these genomes different. Additionally, the KEGG module for nitrogen fixation isn’t comprehensive (as I mentioned before).
So let’s fix that, by using our own set of custom metabolic modules. There are some in the datapack at 00_DATA/modules/. They include a nitrogen fixation module with the full nif gene set (nifHDK and nifENB), a module for hydrogen recycling (hyaABD and hypABCDEF), a module for hopanoid lipid production (squalene synthase, squalene-hopene cyclase, and hpnABGH), and a module for nitrite/nitrate transport (nark and tauABC). The first three are related to nitrogen fixation and associated metabolic activities, while the last one is related to nitrogen assimilation (an alternative to nitrogen fixation) – hence, we expect to find only the last module complete in T. miru and T. nobis.
Here is the custom module for hopanoid production as an example:
ENTRY NIF003
NAME Hopanoid lipid production
DEFINITION (K00801,COG1562) (K06045,COG1657) PF01370.25 PF00535.30 PF01048.24 PF04055.25+PF11946.12
ORTHOLOGY K00801 farnesyl-diphosphate farnesyltransferase [EC:2.5.1.21]
COG1562 phytoene/squalene synthetase
K06045 squalene-hopene/tetraprenyl-beta-curcumene cyclase [EC:5.4.99.17 4.2.1.129]
COG1657 terpene cyclase SqhC
PF01370.25 Hopanoid-associated sugar epimerase HpnA
PF00535.30 Hopene-associated glycosyltransferase HpnB
PF01048.24 Putative hopanoid-associated phosphorylase HpnG
PF04055.25 Hopanoid biosynthesis associated radical SAM protein HpnH (Radical SAM domain)
PF11946.12 Hopanoid biosynthesis associated radical SAM protein HpnH (unknown associated domain)
CLASS User modules; Biosynthesis; Lipid biosynthesis
ANNOTATION_SOURCE K00801 KOfam
K06045 KOfam
COG1562 COG24_FUNCTION
COG1657 COG24_FUNCTION
PF01370.25 Pfam
PF00535.30 Pfam
PF01048.24 Pfam
PF04055.25 Pfam
PF11946.12 Pfam
///
As you can see, it includes enzymes from multiple annotation sources. We’ve already annotated our genomes with all of those functional databases, so we are good to go.
To set up the custom modules into a modules database that we can use with anvi-estimate-metabolism, we need to use the program anvi-setup-user-modules:
anvi-setup-user-modules -u ../00_DATA/
Doing so creates a database at 00_DATA/USER_MODULES.db containing these four modules. We can now give this database to anvi-estimate-metabolism with the -u parameter. We’ll also use the --only-user-modules flag to skip the KEGG module estimation.
Additionally, we will ask the program to compute module copy numbers for us with the --add-copy-number flag. Copy numbers are usually more suitable for metagenomic input rather than individual genomes, but transporter genes can often occur in multiple copies in a single genome and we want to be able to capture that signal in our estimation output.
anvi-estimate-metabolism -e external-genomes.txt \
-u ../00_DATA/ \
--only-user-modules \
--add-copy-number \
-O nitrogen_metabolism
Take a look at the output (nitrogen_metabolism_modules.txt). What do you notice?
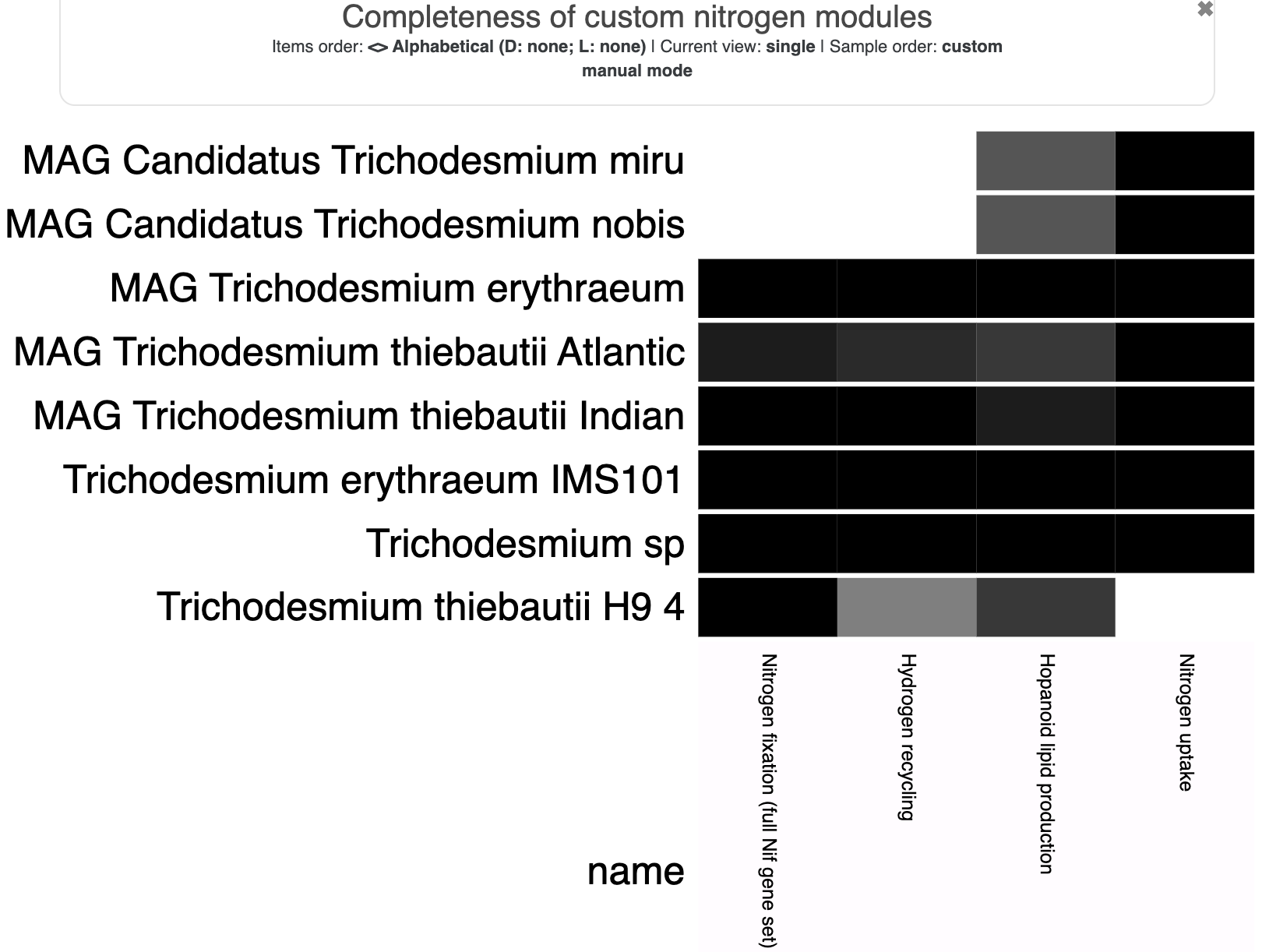
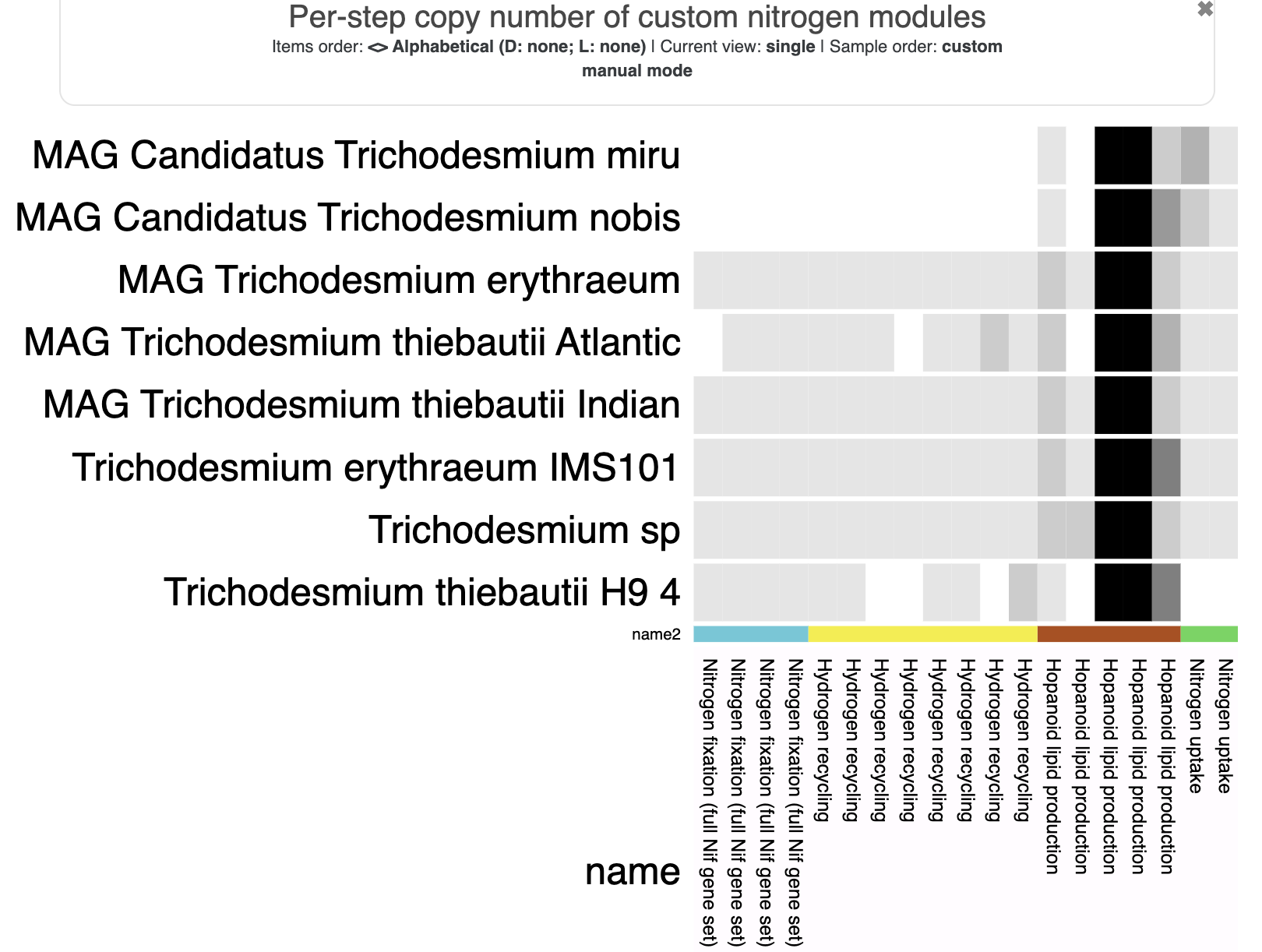
For convenience, I’ll also show the heatmap of pathway completeness scores and the heatmap of per-step copy numbers:
In these visualizations, we’ve adjusted the min/max values to better show the different values. The completeness score heatmap has a minimum of 0.25 (so any completeness value below that appears white) and the normal maximum of 1.0. The copy number heatmap has the normal minimum of 0 and a maximum of 10 (so any copy number above 10 appears black) – there are some steps that have way more than 10 copies, which makes the typical range of 0-2 copies extremely difficult to see on the heatmap unless we cap the value.
Show/Hide Commands to generate the heatmaps
Just like before, if you want a heatmap, you’ll have to generate the output in matrix format:
anvi-estimate-metabolism -e external-genomes.txt \
-u ../00_DATA/ \
--only-user-modules \
--add-copy-number \
-O nitrogen_metabolism \
--matrix-format
For a heatmap this small, we don’t really need to cluster anything. But it would be nice to see the module names instead of just their numbers, so we’ll make a little table of names:
echo -e "module\tname" > custom_mod_names.txt
sqlite3 ../00_DATA/USER_MODULES.db "select module,data_value from modules where data_name='NAME'" | \
tr '|' '\t' >> custom_mod_names.txt
Since we need a profile database before we can store any additional data in it, we’ll go ahead and run anvi-interactive with the --dry-run flag to create one. Then we can import the module name data. If you want, you can also import a state file to automatically make the visualization into a heatmap.
anvi-interactive -d nitrogen_metabolism-module_pathwise_completeness-MATRIX.txt \
-p nitrogen_completeness.db \
--manual \
--title "Completeness of custom nitrogen modules" \
--dry-run
anvi-import-misc-data -p nitrogen_completeness.db -t items custom_mod_names.txt
# if you want, import the heatmap visualization settings
anvi-import-state -p nitrogen_completeness.db -n default -s ../00_DATA/nitrogen_heatmap.json
Finally, here is the visualization command (this time without --dry-run):
anvi-interactive -d nitrogen_metabolism-module_pathwise_completeness-MATRIX.txt \
-p nitrogen_completeness.db \
--manual \
--title "Completeness of custom nitrogen modules"
If you also want to make the per-step copy number heatmap, it is a similar chain of commands. The only difference is the module name info file, which now has to be based on the step names. Here are all the commands to run:
# a little BASH loop to generate the step names file
# with a little trick: two columns for the name so that one can be colors and one can be text
echo -e "step\tname\tname2" > custom_step_names.txt
while read step; do \
mod=$(echo $step | cut -d '_' -f 1); \
name=$(grep $mod custom_mod_names.txt | cut -f 2); \
echo -e "$step\t$name\t$name" >> custom_step_names.txt; \
done < <(cut -f 1 nitrogen_metabolism-step_copy_number-MATRIX.txt | tail -n+2)
# get a profile db
anvi-interactive -d nitrogen_metabolism-step_copy_number-MATRIX.txt \
-p nitrogen_step_copies.db \
--manual \
--title "Per-step copy number of custom nitrogen modules" \
--dry-run
# import relevant data
anvi-import-misc-data -p nitrogen_step_copies.db -t items custom_step_names.txt
anvi-import-state -p nitrogen_step_copies.db -n default -s ../00_DATA/nitrogen_step_copies.json
# visualize
anvi-interactive -d nitrogen_metabolism-step_copy_number-MATRIX.txt \
-p nitrogen_step_copies.db \
--manual \
--title "Per-step copy number of custom nitrogen modules"
Here are some of my observations:
- As we expected, T. miru and T. nobis only have the
NIF004(Nitrogen uptake) module complete. - The other 6 genomes have all the modules >80% complete (except for H9, which is missing several genes from the hydrogen recycling and nitrogen uptake modules. But we already know it is quite an incomplete genome).
- T. miru and T. nobis have multiple copies of the narK transporter (as Tom found in his paper) while the others each have one. This isn’t enough to make the overall nitrogen uptake module have a higher copy number, but you can see the copies of the individual transporters in the last column (
per_step_copy_numbers), and in the copy number heatmap. - Interestingly, the hopanoid lipid production module (
NIF003) has relatively high completeness in most genomes (including T. miru and T. nobis, in which the module is 75% complete), and a lot of that seems to result from finding many Pfam annotations for the hpn gene domains. This contrasts with the results from Tom’s paper – Tom used the RAST annotation tool to find the hpnABGH genes, which may have been a more stringent and/or specific strategy. Perhaps these Pfam domains are too generic to indicate hopanoid production? If we were serious about this analysis, we would probably cross-check our module with a lipid biosynthesis expert to make sure it is appropriate for identifying this metabolic capacity. :) - One thing looks weird! If you look at the per-step copy numbers for
NIF003, the last step always has a copy number of 0 – even though there are certainly genomes in which both hpnH Pfam domains are annotated. In fact, for 6 of the genomes, the pathwise copy number is 1 (or 2) while the stepwise copy number is 0.
Let’s look into that last point a bit more. What is going on with the stepwise copy number estimation for hopanoid production? When things look weird in the results, it is always good to take a single genome, rerun the metabolism estimation, and pay attention to the warnings in the terminal output. Terminal output is more verbose on individual genomes than in ‘multi-mode’ for anvi-estimate-metabolism.
The T. erythraeum genome shows this weird copy number pattern, so let’s use that one:
anvi-estimate-metabolism -c ../Trichodesmium_erythraeum_IMS101-contigs.db \
-u ../00_DATA/ \
--only-user-modules \
-O test \
--add-copy-number
Aha! There is a relevant warning in the terminal output:
WARNING
===============================================
The gene call 208 has multiple annotations to alternative enzymes within the
same step of a metabolic pathway (PF04055.25, PF11946.12), and these enzymes
unfortunately have a complex relationship. The affected module is NIF003, and
here is the step in question: PF04055.25+PF11946.12. We arbitrarily kept only
one of the annotations to this gene in order to avoid inflating the step's copy
number, but due to the complex relationship between these alternatives, this
could mean that the copy number for this step is actually too low. Please heed
this warning and double check the stepwise copy number results for NIF003 and
other pathways containing gene call 208.
The lessons here: pay attention to warnings from anvi’o programs. And keep in mind that many of these warnings are suppressed when processing multiple inputs, so testing things on individual genomes might be the way to go when outputs look strange.
If something goes wrong (or weird) while using anvi’o, you may want to try re-running whatever you just did with the addition of the global --debug flag (which works for all anvi’o programs). This flag enables extra terminal output, which in the best case may help you figure out what is going on, and in the worst case can provide enough information to send to the developers when you ask them for help (in case of errors, we especially appreciate the code tracebacks that --debug allows you to see).
If you found this section useful and you want to make your own custom metabolic modules, check out this guide on the user-modules-data help page.
Reaction networks and drawing KEGG Pathway Maps
Let’s move onto the second type of metabolism reconstruction: metabolic modeling. Anvi’o can generate a reaction-network from the KEGG Ortholog (KO) annotations in any contigs database or pangenome database. The network connects all genes with KO annotations to the chemical reactions they catalyze, and the metabolites consumed or produced by those reactions. Reaction and compound information are taken from the ModelSEED database.
If you want to use these programs, you will first have to run anvi-setup-modelseed-database (if you haven’t already done so in your anvi’o environment).
Let’s make a reaction network for one genome using the program anvi-reaction-network:
anvi-reaction-network -c ../Trichodesmium_sp-contigs.db
There will be plenty of output on your terminal screen, but no output files added to your working directory – the network will be stored directly in the contigs database.
Now, if you wanted to do some flux balance analysis (FBA) to model the flow of metabolites through this network – bad news, you can’t do that in anvi’o. However, you can export the reaction-network into a JSON file suitable for common metabolic modeling software (like COBRApy) with the program anvi-get-metabolic-model-file:
anvi-get-metabolic-model-file -c ../Trichodesmium_sp-contigs.db \
-o Trichodesmium_sp_rxn_network.json
The output file is really big, because it contains every single metabolite, reaction, and gene contributing to the reaction network.
What we will do in anvi’o is to use this reaction-network for some neat visualizations of KEGG Pathway Maps with the program anvi-draw-kegg-pathways.
anvi-draw-kegg-pathways --contigs-dbs ../Trichodesmium_sp-contigs.db \
-o Trichodesmium_sp_PATHWAY_MAPS \
--ko
For each KEGG Pathway Map, the program will highlight the KOs from the map that are annotated in each provided genome, and create a PDF file in the specified output directory. We only provided a single genome, so the resulting maps are specific to that genome’s annotations.
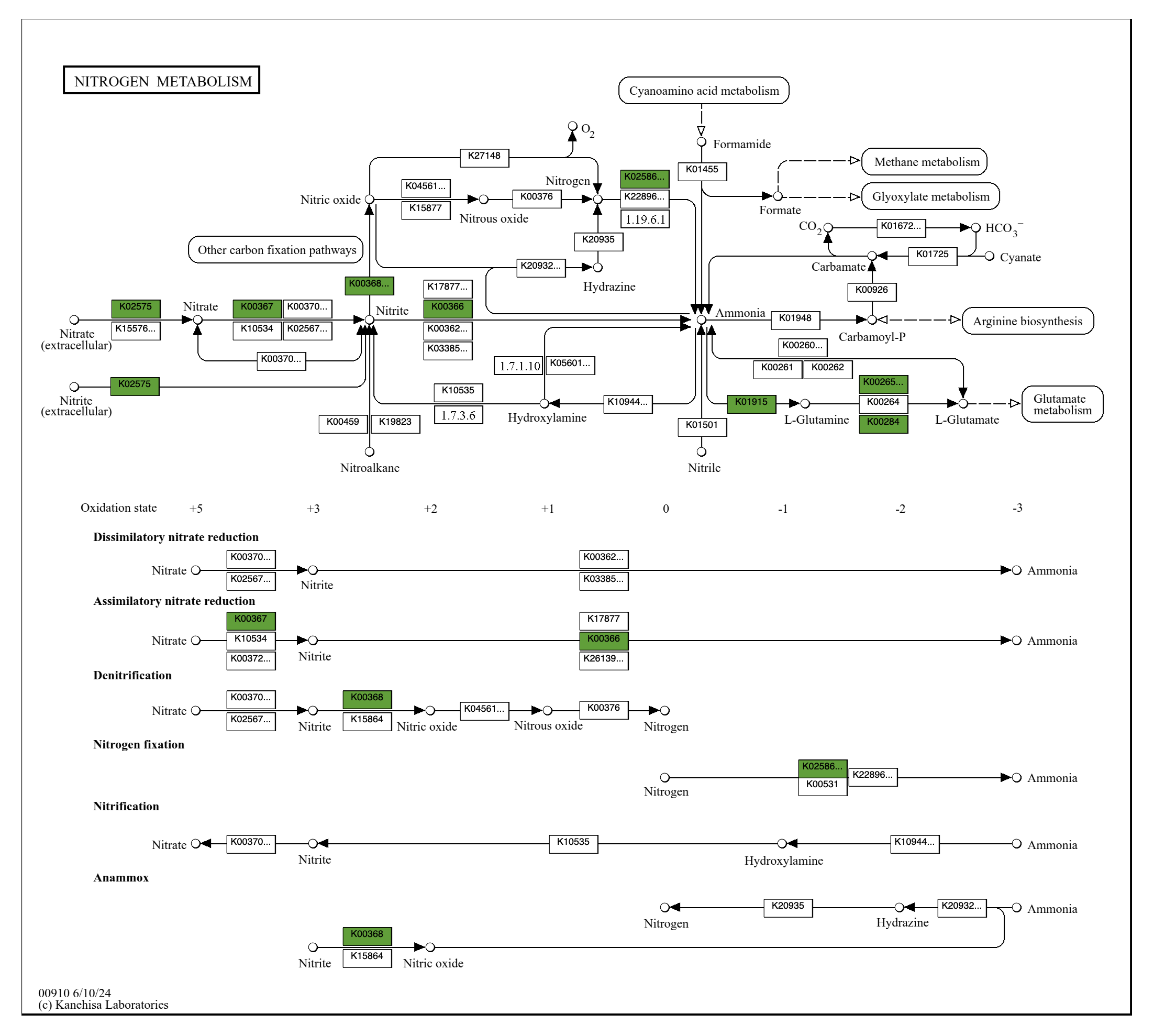
Let’s look at an example map. Of course we will look at the Nitrogen Metabolism map (which is map 00910 and stored at Trichodesmium_sp_PATHWAY_MAPS/kos_00910.pdf):
You can see that in addition to nitrogen fixation (nitrogen to ammonia), this microbe can also import nitrate and nitrite from the extracellular matrix into the cell, convert nitrate to nitrite (assimilatory nitrate reduction) and to nitric oxide, convert nitrite to ammonia, and feed that ammonia into glutamate metabolism – which is the start of amino acid biosynthesis using that fresh new bioavailable nitrogen.
While pathway prediction gives you quantitative summaries of metabolic capacity, these Pathway Map drawings are great for understanding what is actually going on in those pathways, and for seeing the connections between different classes of metabolism. Each map gives you an overall picture of a small-ish, digestible and ecologically-relevant part of the entire reaction network.
I bet you are wondering how this map looks different across our Trichodesmium genomes. If so, then you are in luck, because we can also use anvi-draw-kegg-pathways to compare metabolic capacity of multiple genomes.
First, we will need to run anvi-reaction-network on all the other genomes. This program unfortunately doesn’t accept an external genomes file as input; however, we can reuse our BASH loop strategy from chapter 1.
while read genome
do
anvi-reaction-network -c ../${genome}-contigs.db; \
done < ../genomes.txt
Once that is done, we can run anvi-draw-kegg-pathways again, and this time we can provide an external genomes file. We will also provide the --draw-grid flag so that we get maps showing the results from each individual genome (in addition to the default map showing the total annotations across all genomes). Since it would take a long time to draw every single Pathway Map for all 8 genomes, we will focus on a subset of maps as specified by the --pathway-numbers parameter – the list below includes a couple of maps related to nitrogen metabolism, a few for photosynthesis and related metabolic capacities, and a map for metabolism of some amino acids.
anvi-draw-kegg-pathways --external-genomes ../external-genomes.txt \
-o ALL_PATHWAY_MAPS \
--ko \
--draw-grid \
--pathway-numbers 00910 01310 00195 00860 00906 00900 00260 00290
The resulting files in the ALL_PATHWAY_MAPS include:
- multiple map
.pdffiles (one per requested Pathway Map) with colored boxes showing the number of annotations to a given KO across all input genomes - a
colorbar.pdfshowing the legend – what color corresponds to what count - a
grid/subdirectory containing similarly-named map.pdffiles, except this time each one shows a grid of individual maps for each genome
For consistency with above, we’ll look at Pathway Map 00910 for Nitrogen Metabolism again. Here is the combined map at ALL_PATHWAY_MAPS/kos_00910.pdf:
The legend tell us that smaller counts are warm colors (starting from a count of 1 in yellow) and higher counts are warm colors (going to a max of 8 in dark purple). So we can tell from this map that all the genomes encode enzymes for assimilatory nitrate reduction, 6 out of the 8 genomes encode nitrogen fixation, and only 1 genome can convert nitrite to ammonia. But we don’t know which genome can do what.
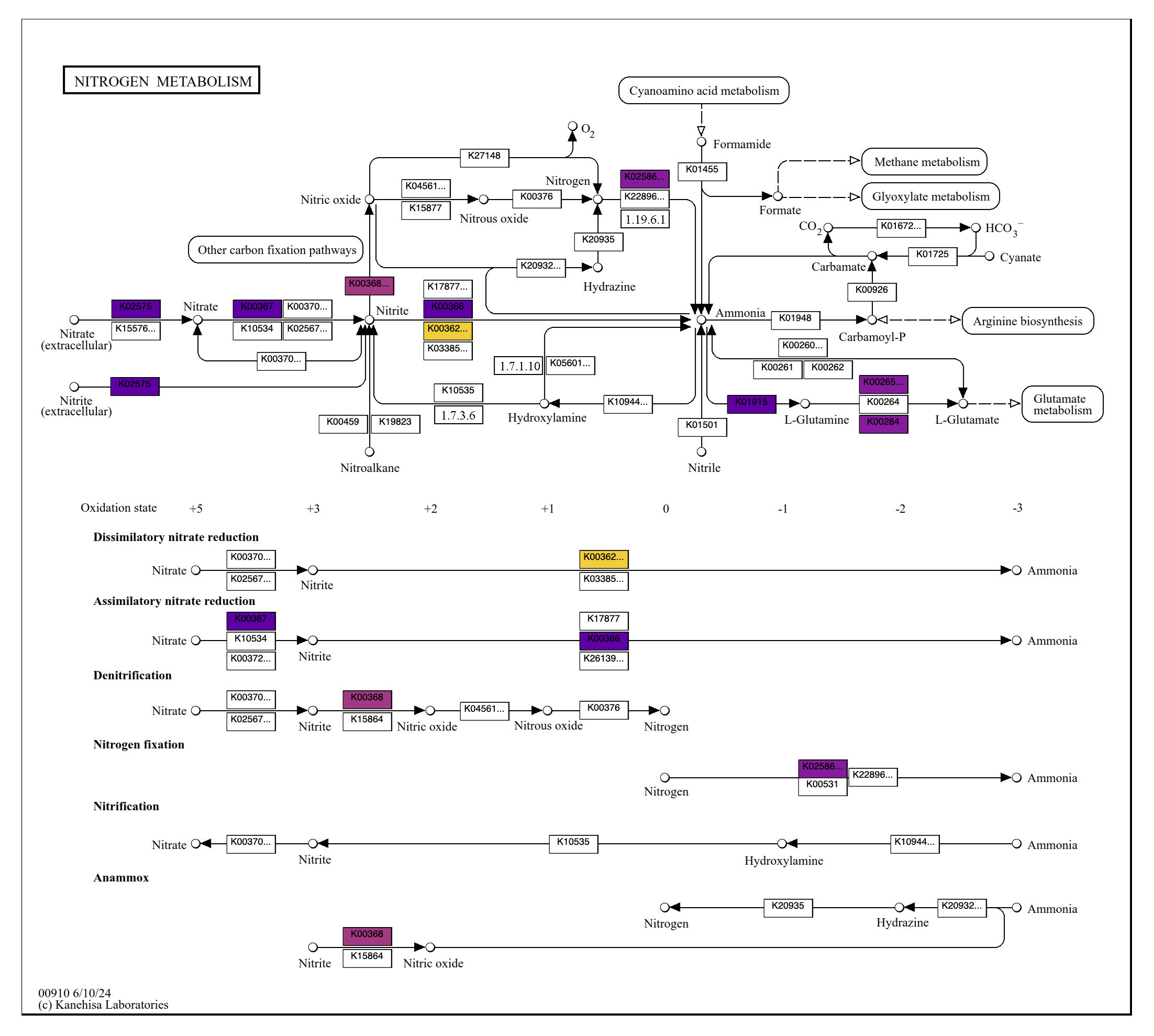
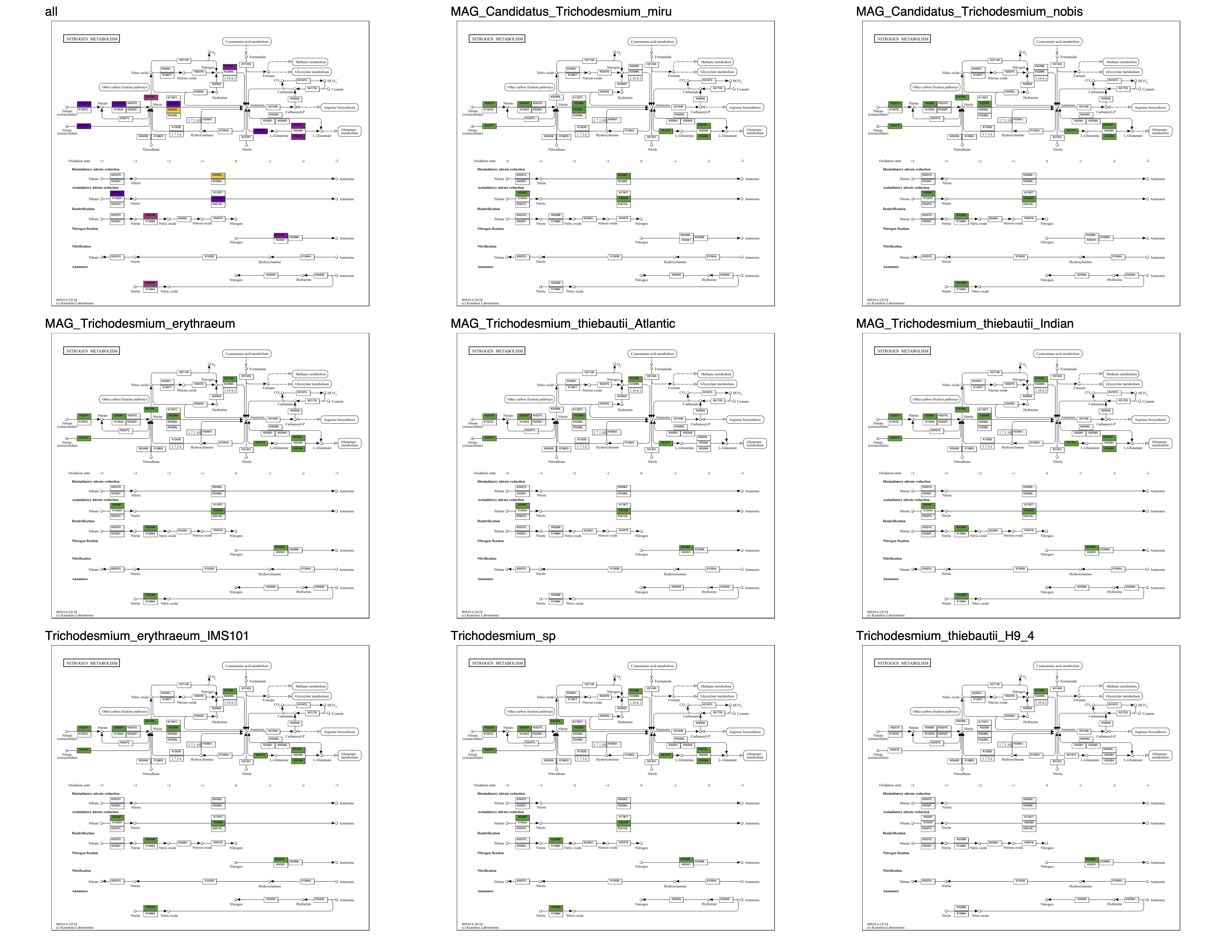
To reveal the specific distribution of enzymes across genomes, here is the corresponding grid map (ALL_PATHWAY_MAPS/grid/kos_00910.pdf):
Now it is clear that T. miru and T. nobis are missing nitrogen fixation (as we know), and that T. miru is the only species that can convert nitrite to ammonia.
I encourage you to look through the other maps (both overall and in grid form). Here are some things to look out for:
- The nitrogen cycle map (01310) shows metabolic capacity with colored reaction arrows rather than boxes. There again you can see that T. miru is the only one doing nitrogen dissimilation.
- In map 00260 (Glycine, Serine and Threonine Metabolism), you can see that T. miru and T. nobis are also the only genomes with the enzyme to convert threonine to 2-oxobutanoate (a precursor leading into the production of several other amino acids). You can see the same thing in map 00290 (Valine, Leucine and Isoleucine Biosynthesis). Clearly the other genomes have the capacity to make those amino acids, just not by starting from 2-oxobutanoate. So biosynthesis of amino acids is just a bit more connected in T. miru and T. nobis compared to the others.
- Map 00195 shows photosynthesis. Maps 00860, 00900, and 00906 are photosynthesis-related capacities, like the biosynthesis of pigments. These are things that all the Trichodesmium species can do, but there are still differences between these genomes – some genomes are missing enzymes that all the others have (a detection issue?) while others have some enzymes that all the others don’t (interesting ecology?).
The KEGG Pathway Maps cover many metabolic capacities that KEGG modules do not, and can be a great tool for comparative metabolism analyses.
Predicting metabolic interactions
One cool thing about Trichodesmium is that they have friends. Bacterial friends, to be specific. Trichodesmium colonies harbor associated bacteria like Alteromonas, Roseibium, and Balneolales species. Based on a recent study about a Red Sea T. thiebautii colony and its bacterial associates Koedooder et al 2023, these microbes have a symbiotic relationship – they help each other by exchanging important molecules. The bacterial associates are known to produce siderophores, which help with harvesting iron from dust in their environment and which Trichodesmium cannot make itself. Meanwhile, Trichodesmium can make several B vitamins that the bacterial associates are auxotrophic for.
Wouldn’t it be cool if we could investigate metabolic interactions using our genomes, without having to painstakingly search for individual gene annotations? Anvi’o got u, fam.
There is a program called anvi-predict-metabolic-exchanges that leverages the reaction-network infrastructure to identify potential metabolites that could be exchanged between two microbes from their genomic data. It does so by examining the reactions in the network to find metabolites that could be produced by only one of the organisms and consumed by the other. If possible, it then leverages the KEGG Pathway Maps that we saw in the previous section to walk over targeted subsets of the network starting from the potential exchange point, in order to compute statistics about the reaction chains leading up to (or away from) the exchange. Those statistics – length, amount of overlap between genomes, etc – serve as evidence for the likelihood of a potential exchange. The program also identifies which compounds are uniquely produced or consumed by one of the genomes in the pair. You can find more technical information about how anvi-predict-metabolic-exchanges works here.
anvi-predict-metabolic-exchanges is an extremely new program as of October 2025. It is not fully refined yet and there may be some remaining bugs. Please validate the results carefully and report issues if you notice weird results. The developers thank you for your patience :)
We are going to test this program out on our T. thiebautii (Atlantic) genome and three genomes of known associated bacteria based on the Koedooder et al 2023 study. The MAGs from that study are not yet publicly available, so instead we will work with reference genomes – one for Alteromonas macleodii (GCF_002849875.1), one for Roseibium aggregatum (GCF_000168975.1), and one for Marinobacter salarius (GCF_002116735.1).
You’ve already learned how to do all the preparatory steps for analyzing genomes – making a contigs database, annotating with KOfams, and creating the reaction network. Feel free to practice on these three new genomes. But in case you just want to get right to the metabolic interactions analysis, there are pre-made, annotated contigs databases for these genomes in the datapack at 00_DATA/associate_dbs/.
You can copy the databases over to your current working directory (assuming you are still in the 02_METABOLISM folder) like so:
cp ../00_DATA/associate_dbs/*.db .
For those interested, click the Show/Hide box below to see how we made those databases.
Show/Hide Steps to make the contigs databases
# download and reformat
curl https://ftp.ncbi.nlm.nih.gov/genomes/all/GCF/002/849/875/GCF_002849875.1_ASM284987v1/GCF_002849875.1_ASM284987v1_genomic.fna.gz -o GCF_002849875.1_ASM284987v1_genomic.fna.gz
gunzip GCF_002849875.1_ASM284987v1_genomic.fna.gz
anvi-script-reformat-fasta GCF_002849875.1_ASM284987v1_genomic.fna -o A_macleodii.fasta --simplify-names --prefix GCF_002849875 --seq-type NT
curl https://ftp.ncbi.nlm.nih.gov/genomes/all/GCF/000/168/975/GCF_000168975.1_ASM16897v1/GCF_000168975.1_ASM16897v1_genomic.fna.gz -o GCF_000168975.1_ASM16897v1_genomic.fna.gz
gunzip GCF_000168975.1_ASM16897v1_genomic.fna.gz
anvi-script-reformat-fasta GCF_000168975.1_ASM16897v1_genomic.fna -o R_aggregatum.fasta --simplify-names --prefix GCF_000168975 --seq-type NT
curl https://ftp.ncbi.nlm.nih.gov/genomes/all/GCF/002/116/735/GCF_002116735.1_ASM211673v1/GCF_002116735.1_ASM211673v1_genomic.fna.gz -o GCF_002116735.1_ASM211673v1_genomic.fna.gz
gunzip GCF_002116735.1_ASM211673v1_genomic.fna.gz
anvi-script-reformat-fasta GCF_002116735.1_ASM211673v1_genomic.fna -o M_salarius.fasta --simplify-names --prefix GCF_002116735 --seq-type NT
rm GCF_00*.fna
# make contigs databases
for g in A_macleodii R_aggregatum M_salarius; do
anvi-gen-contigs-database -f $g.fasta -o $g-contigs.db -T 3
done
# annotate with KOfams
for g in A_macleodii R_aggregatum M_salarius; do
anvi-run-kegg-kofams -c $g-contigs.db -T 4;
done
# make reaction network
for g in A_macleodii R_aggregatum M_salarius; do
anvi-reaction-network -c $g-contigs.db;
done
Since the databases have reaction networks already stored inside them, we can get right to predicting interactions. Let’s start by running anvi-predict-metabolic-exchanges on one pair of genomes:
anvi-predict-metabolic-exchanges -c1 ../MAG_Trichodesmium_thiebautii_Atlantic-contigs.db \
-c2 A_macleodii-contigs.db \
-O thiebautii-vs-macleodii \
-T 4
There were probably some warnings on your screen about some Pathway Maps not having a “reaction (RN) type KGML file”. This just means that we couldn’t use those specific Maps when doing the Pathway Map walks for computing evidence. If you don’t want to see the warnings, you can specifically exclude the missing Maps with the flag --exclude-pathway-maps (we will do this later).
There are actually two prediction strategies used by anvi-predict-metabolic-exchanges, and the terminal output will tell you how many predictions were made using each one. Briefly, the first strategy (“Pathway Map Walk”) is based on the KEGG Pathway Maps – it works on only a subset of metabolites, but it generates high-quality predictions that are associated with evidence from the Pathway Map walks. The second strategy (“Reaction Network Subset”) works on all metabolites in the network (even those not in KEGG Pathway Maps), but generates predictions that are potentially less accurate. That distinction is reflected in the sheer number of predictions that we get from either approach:
Number of exchanged compounds predicted from KEGG Pathway Map walks : 128
Number of unique compounds predicted from KEGG Pathway Map walks : 621
(...)
Number of exchanged compounds predicted from Reaction Network subset approach : 315
Number of unique compounds predicted from Reaction Network subset approach : 1,297
The Pathway Map walk approach gives you fewer predictions, while the reaction network subset approach gives you more (but lower confidence) predictions. Note that you can turn off either approach by using the --no-pathway-walk or --pathway-walk-only flags, respectively. When you use both approaches at once, the network subset strategy only works on compounds that have not yet been processed as part of the Pathway Map walk strategy.
Here is the summary of the results from the terminal:
OVERALL RESULTS
===============================================
Identified 443 potentially exchanged compounds and 1918 compounds unique to one
genome.
These results are described in three output files: thiebautii-vs-macleodii-potentially-exchanged-compounds.txt describes the potential exchanges (with summaries of the evidence from associated Pathway Maps), thiebautii-vs-macleodii-evidence.txt contains the full set of statistics from all Pathway Map walks, and thiebautii-vs-macleodii-unique-compounds.txt describes the metabolites that are uniquely produced or consumed by one of the organisms. All of these output files are indexed by ModelSEED compound ID numbers.
Our predictions are limited to the metabolites we can find in the ModelSEED database. ModelSEED doesn’t contain compounds for siderophores like petrobactin, vibrioferrin, or agrobactin (at least, it didn’t at the time of writing this tutorial). So we cannot find those specific interactions as described in the Koedooder et al 2023 paper.
But we might be able to find the B vitamin exchanges mentioned in that paper, or perhaps something new! Let’s take a careful look at the predicted exchanges. Here are the first 10 lines of that file:
compound_id |
compound_name |
genomes |
produced_by |
consumed_by |
prediction_method |
max_reaction_chain_length |
max_production_chain_length |
production_overlap_length |
production_overlap_proportion |
production_chain_pathway_map |
max_consumption_chain_length |
consumption_overlap_length |
consumption_overlap_proportion |
consumption_chain_pathway_map |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| cpd00027 | D-Glucose | A_macleodii,MAG_Trichodesmium_thiebautii_Atlantic | A_macleodii | A_macleodii,MAG_Trichodesmium_thiebautii_Atlantic | Pathway_Map_Walk | 10 | 8 | None | None | 00500 | 2 | 2 | 1.0 | 00500 |
| cpd00037 | UDP-N-acetylglucosamine | A_macleodii,MAG_Trichodesmium_thiebautii_Atlantic | A_macleodii | A_macleodii,MAG_Trichodesmium_thiebautii_Atlantic | Pathway_Map_Walk | 14 | 6 | None | None | 00520 | 8 | 8 | 1.0 | 00550 |
| cpd00044 | 3-phosphoadenylylsulfate | A_macleodii,MAG_Trichodesmium_thiebautii_Atlantic | A_macleodii | A_macleodii,MAG_Trichodesmium_thiebautii_Atlantic | Pathway_Map_Walk | 9 | 6 | None | None | 00920 | 3 | 3 | 1.0 | 00920 |
| cpd00065 | L-Tryptophan | A_macleodii,MAG_Trichodesmium_thiebautii_Atlantic | A_macleodii | A_macleodii,MAG_Trichodesmium_thiebautii_Atlantic | Pathway_Map_Walk | 14 | 13 | None | None | 00400 | 1 | 1 | 1.0 | 00380 |
| cpd00098 | Choline | A_macleodii,MAG_Trichodesmium_thiebautii_Atlantic | A_macleodii,MAG_Trichodesmium_thiebautii_Atlantic | A_macleodii | Pathway_Map_Walk | 3 | 1 | 1 | 1.0 | 00564 | 2 | None | None | 00260,00670 |
| cpd00100 | Glycerol | A_macleodii,MAG_Trichodesmium_thiebautii_Atlantic | A_macleodii | A_macleodii,MAG_Trichodesmium_thiebautii_Atlantic | Pathway_Map_Walk | 2 | 1 | None | None | 00052,00561 | 1 | 1 | 1.0 | 00561 |
| cpd00104 | BIOT | A_macleodii,MAG_Trichodesmium_thiebautii_Atlantic | A_macleodii,MAG_Trichodesmium_thiebautii_Atlantic | A_macleodii | Pathway_Map_Walk | 5 | 2 | 2 | 1.0 | 00780 | 3 | None | None | 00780 |
| cpd00107 | L-Leucine | A_macleodii,MAG_Trichodesmium_thiebautii_Atlantic | A_macleodii | A_macleodii,MAG_Trichodesmium_thiebautii_Atlantic | Pathway_Map_Walk | 11 | 10 | None | None | 00290 | 1 | 1 | 1.0 | 00970 |
| cpd00108 | Galactose | A_macleodii,MAG_Trichodesmium_thiebautii_Atlantic | A_macleodii | A_macleodii,MAG_Trichodesmium_thiebautii_Atlantic | Pathway_Map_Walk | 8 | 7 | None | None | 00052 | 1 | 1 | 1.0 | 00052 |
The predictions are ranked in order from best evidence to no evidence, so all of these at the top are coming from the Pathway_Map_Walk prediction strategy. And here already is one of our expected exchanges: the compound called ‘BIOT’ is actually biotin aka Vitamin B7, and T. thiebautii is potentially cross-feeding it to A. macleodii as suggested in the Koedooder et al 2023 paper.
There are also a few unexpected predictions that look interesting: A. macleodii seems to be able to produce D-glucose, L-Tryptophan, and L-Leucine while T. thiebautii only can consume these compounds. The relatively long production chain lengths indicate that these aren’t isolated production reactions in the network; they are rather part of a longer metabolic pathway. Meanwhile, the None values for the production overlap mean that the T. thiebautii genome doesn’t contain any of the enzymes in the production-associated Pathway Map. In the case of L-Tryptophan, that map is 00400. Let’s check the associated evidence for L-Tryptophan in map 00400 in the evidence file:
compound |
compound_name |
longest_chain_compound_names |
longest_chain_compounds |
longest_chain_reactions |
longest_reaction_chain_length |
maximum_overlap |
organism |
pathway_map |
proportion_overlap |
type |
|---|---|---|---|---|---|---|---|---|---|---|
| cpd00065 | L-Tryptophan | L-Tryptophan,indol,Indoleglycerol phosphate,1-(2-carboxyphenylamino)-1-deoxyribulose 5-phosphate,N-5-phosphoribosyl-anthranilate,Anthranilate,Chorismate,5-O–1-Carboxyvinyl-3-phosphoshikimate,3-phosphoshikimate,Shikimate,3-Dehydroshikimate,5-Dehydroquinate,DAHP,Phosphoenolpyruvate | C00078,C00463,C03506,C01302,C04302,C00108,C00251,C01269,C03175,C00493,C02637,C00944,C04691,C00074 | rn:R00674,rn:R02340,rn:R03508,rn:R03509,rn:R01073,rn:R00985 rn:R00986,rn:R01714,rn:R03460,rn:R02412,rn:R02413,rn:R03084,rn:R03083,rn:R01826 | 13 | None | A_macleodii | 00400 | None | production |
| cpd00065 | L-Tryptophan | None | None | None | None | None | MAG_Trichodesmium_thiebautii_Atlantic | 00400 | None | consumption |
Show/Hide How to get this table
To get the two rows of information shown above, you can search for L-Tryptophan’s compound ID, followed by a search for the Pathway Map 00400:
grep cpd00065 thiebautii-vs-macleodii-evidence.txt | grep 00400
Okay, so far it looks pretty promising. There is a very long chain of reactions to produce tryptophan from phosphoenolpyruvate in A. macleodii, and nothing at all in T. thiebautii. But just to be sure, let’s also visualize map 00400 for these two genomes with anvi-draw-kegg-pathways (and while we are at it, let’s include the production maps for the other two interesting predictions):
anvi-draw-kegg-pathways --contigs-dbs ../MAG_Trichodesmium_thiebautii_Atlantic-contigs.db A_macleodii-contigs.db \
-o thiebautii-vs-macleodii_PATHWAY_MAPS \
--pathway-numbers 00400 00500 00290 \
--draw-grid \
--ko
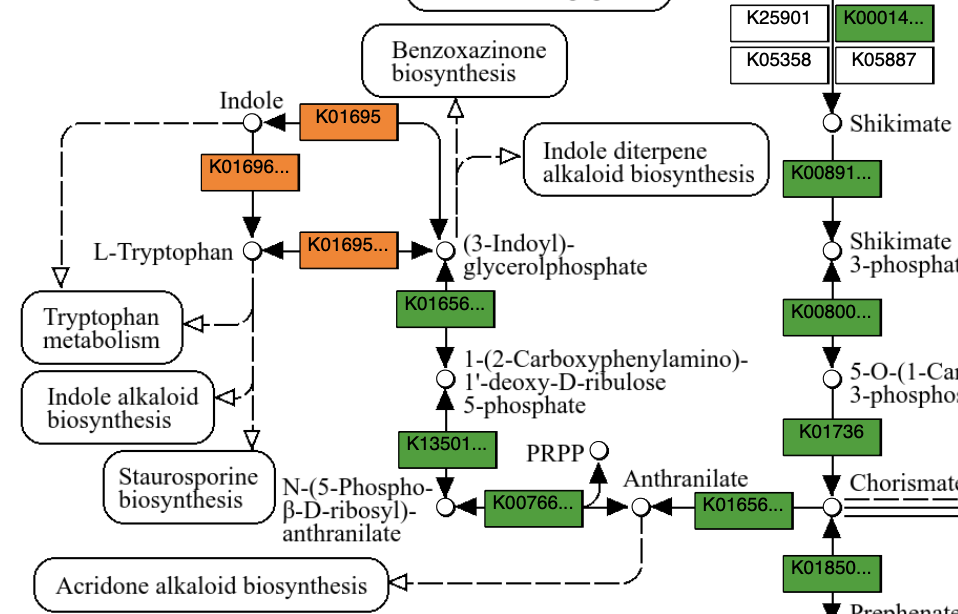
Here is the part of that map where tryptophan is produced:
Oh, wait. Only one of the two genomes (A. macleodii) has the enzymes to convert between indole or indoyl-containing compounds and tryptophan, but clearly both genomes have all the rest of the enzymes leading up to that point (we didn’t detect this in T. thiebautii because the start of the reaction chain was missing in its genome). This could be a legitimate gap in T. thiebautii’s reaction network. Or, we could just be missing the annotation for these functions in T. thiebautii (i.e., either we are missing the relevant chunk of the genome, or we failed to annotate the corresponding gene that is present in the MAG).
Perhaps this is a valid exchange, and perhaps it is not. It would take a bit more digging through the T. thiebautii MAG to see if we could find the relevant enzymes (e.g., K01695) with a more careful homology search. But proving that an enzyme is truly missing is hard, and we don’t necessarily want to do that right now. So we are just going to move on.
I encourage you to take a look at the corresponding maps for the production of L-Leucine and D-Glucose. The L-Leucine situation (in map 00290) looks very similar to the L-Tryptophan situation – most of the enzymes are there in both genomes, but T. thiebautii is missing just the tail end of the reaction chain such that it looks like this microbe cannot fully synthesize several amino acids, including leucine, valine and isoleucine. Based on our previous exploration of this pathway map across all of the Trichodesmium genomes, this seems unlikely to be the true story. Most of the other genomes have those enzymes, including the T. thiebautii MAG from the Indian Ocean. So we can ignore this prediction.
For D-Glucose (map 00500), on the other hand, there are many enzymes in the reaction chain that A. macleodii has but T. thiebautii does not. So that prediction looks legitimate – A. macleodii could be cross-feeding glucose to T. thiebautii. Whether or not that actually happens in real life is uncertain. First, A. macleodii would have to intentially create glucose (as opposed to funneling its energy and carbon into other things). Second, it would have to produce more glucose than it individually needs, and third, that glucose would have to make its way outside of the cell where T. thiebautii could pick it up. But this potential exchange of glucose is now a hypothesis that is testable either by experimental means or literature review.
Regardless, one lesson to take away from this section is: predicting metabolic exchanges is fallible, so you should always check the predictions carefully before jumping to conclusions. Drawing the KEGG Pathway Maps is a great visual tool to help with this.
Allowing for gaps in reaction chains
You might be wondering if there is a way to change the stringency of the Pathway Map walks so that they can handle small gaps. Indeed there is! We can set the --maximum-gaps parameter to allow for some number of gaps in the reaction chains. Unfortunately, this won’t fix the case of L-Tryptophan or L-Leucine, because the gap occurs right at the start of the reaction chain and you cannot build a chain that starts from nothing (Iva and Sam need to think about how to get around this edge case). However, it could help create a better ranking for other ambiguous cases in which the gaps are internal to the reaction chain.
So let’s run the predictions again – this time allowing for a gap of 1 missing enzyme. We will also exclude those problematic pathway maps that gave us pesky warnings before. While creating the tutorial, I noticed that allowing for a gap of 1 causes problems with the Pathway Walk on a specific map, 00061 (Fatty acid biosynthesis) – processing that map takes forever, potentially because the gap causes a cycle that the codebase can’t handle quite yet. To avoid the long (potentially infinite) runtime, we will also exclude this specific map.
How did I figure out that map 00061 was the problematic one? I re-ran the program with the --debug flag to get more verbose output, and noticed that map 00061 was the only one that started processing but didn’t finish.
In addition, since we’ve been seeing amino acids in the prediction output, we will add the --use-equivalent-amino-acids flag to make sure that we are considering metabolites like “Lysine” and “L-Lysine” to be the same (which could affect our results). Here is the full command:
anvi-predict-metabolic-exchanges -c1 ../MAG_Trichodesmium_thiebautii_Atlantic-contigs.db \
-c2 A_macleodii-contigs.db \
-O thiebautii-vs-macleodii-gap1 \
--maximum-gaps 1 \
--use-equivalent-amino-acids \
--exclude-pathway-maps 00061,00121,00190,00195,00196,00511,00542,00543 \
-T 4
This time, the number of predictions from the Pathway Map walk approach is a little bit smaller (122 predictions, compared to 128 from before). Some predictions likely went away because we excluded an additional Pathway Map. But the top-ranked results in the table above mostly didn’t change, except for some of the reaction chains getting a bit longer.
So now is the time to do some targeted searches through the rest of the file. You can look through the rest of the predictions using whatever strategy you think is best. I would recommend focusing on our first set of output files (the ones made without considering gaps), as those are more stringent and seem to be more reliable predictions overall.
Confused by the ModelSEED compound names? You are not a biochemist and are overwhelmed with all this molecular information? Us, too. One option for you is to take the output, pick out the predictions that you have high-confidence in, and give that list to a large-language model (LLM) so it can pick out the ones that are likely to be biologically-relevant in your system. Then you can focus your efforts on carefully validating those predictions.
Here are some of the interesting predictions that I found:
- Nitrate (cpd00209), produced by A. macleodii and consumed by T. thiebautii. You might remember from our earlier exploration of Pathway Map 00910 that T. thiebautii has the enzymes to import extracellular nitrate and convert it to nitrite. Perhaps some of that exterior nitrate is coming from its associated bacteria.
- Thymine (cpd00151) or Thymidine (cpd00184), produced by T. thiebautii and consumed by A. macleodii. These are part of ‘T’ nucleotides and are needed by both organisms. Other marine microbes are known to exchange nucleotide components, so perhaps these do, too.
Analyzing multiple pairs of genomes
When we are working with a whole consortium of microbes, we can ask anvi-predict-metabolic-exchanges to process all genomes by providing an external genomes file. If you don’t do anything else, the program will process all possible pairs of genomes. But you can also provide a genome-pairs file to specify which pairwise comparisons it should do.
In our case, we have four genomes – T. thiebautii and three associated bacteria – and we mostly care about the interactions between T. thiebautii and its associates. We’ve also already analyzed T. thiebautii vs A. macleodii in the previous section. So we’ll provide a genome-pairs file indicating only T. thiebautii vs R. aggregatum and T. thiebautii vs M. salarius. That file is in your datapack already, and here is what it looks like:
genome_1 |
genome_2 |
|---|---|
| MAG_Trichodesmium_thiebautii_Atlantic | R_aggregatum |
| MAG_Trichodesmium_thiebautii_Atlantic | M_salarius |
You can copy this file over to your current working directory:
cp ../00_DATA/genome-pairs.txt .
The names in the genome-pairs file must match the corresponding external-genomes file. We can make an external genomes file for our 3 associated bacteria, whose contigs-dbs are in our working directory. And we can append to that the line for the T. thiebautii (Atlantic) MAG from our previous external genomes file:
anvi-script-gen-genomes-file --input-dir . -o consortium_external_genomes.txt
grep thiebautii_Atlantic ../external-genomes.txt >> consortium_external_genomes.txt
Here is how you can run anvi-predict-metabolic-exchanges on those pairs:
anvi-predict-metabolic-exchanges -e consortium_external_genomes.txt \
--genome-pairs-txt genome-pairs.txt \
-O thiebautii-vs-many \
--use-equivalent-amino-acids \
--exclude-pathway-maps 00061,00071,00121,00190,00195,00196,00270,00511,00542,00543 \
-T 2 \
-P 2
We are excluding several pathway maps here because our algorithms cannot currently handle them (or cannot process them efficiently enough for a tutorial setting).
Notice that here we set the number of processes (-P) to 2, so that two genome pairs can be processed in parallel. And we set the number of threads per process (-T) to 2 as well, so in total the program will be using 4 threads.
When the program finishes, you will get similar output files as before, except that multiple pairs of genomes will be included in the results. The predictions include many of the same exchanges that we saw between T. thiebautii and A. macleodii before, like ‘BIOT’, nitrate, and thymine. So it seems like the bacterial associates have similar interactions with their cyanobacterial partner.
Metabolic conclusions
We’ve now used several different strategies to investigate the metabolism of our Trichodesmium genomes and their bacterial associates. We’ve once again recapitulated the lack of nitrogen fixation in T. miru and T. nobis, and we’ve tried some very new programs for visualizing KEGG Pathway Maps and for predicting metabolic interactions.
Don’t forget to go back to the parent directory before you move on to the next tutorial section:
cd ..
Congrats!
This is currently the last chapter of the tutorial. Well done! We hope it was useful for you.
If you want to go back to the main page of the tutorial, click here.
If you have any questions about this exercise, or have ideas to make it better, please feel free to get in touch with the anvi’o community through our Discord server: