An anvi'o tutorial with Trichodesmium genomes (Chapter 1)
Table of Contents
- Quick Navigation
- Chapter 1 - Genomics
- Download and reformat the genome
- Generate a contigs database
- Annotate single-copy core genes and Ribosomal RNAs
- Estimate completeness and redundancy
- Estimate SCG taxonomy
- Functional annotations
- Working with multiple genomes
- Working with one (or more) metagenomes
- The next chapter
About this page This webpage is one chapter of a much larger effort to cover multiple aspects of anvi’o in the same tutorial. If you need more context, please visit the main page of the tutorial, where you will find information about the dataset we are working with and the commands to download the tutorial datapack.
Quick Navigation
- Tutorial introduction (main page)
- Chapter 1: Genomics ← you are here
- Chapter 2: Pangenomics
- Chapter 3: Phylogenomics
- Chapter 4: Metabolism
Chapter 1 - Genomics
To introduce you into the anvi’o-verse, we will run some basic genomics analysis on a single genome. In this case, we will download a known Trichodesmium genome from NCBI to add to our collection. We will learn how to:
- reformat a FASTA file
- generate a contigs-db
- annotate gene calls with single-copy core genes and functions
- estimate the taxonomy and completeness/redundancy of the genome
- get data out of the database and into parseable text files
Let’s get started.
Download and reformat the genome
We will supplement our set of Trichodesmium genomes with this Trichodesmium MAG from NCBI. It is labeled as Trichodesmium sp. MAG_R01, so we don’t know exactly what species of Trichodesmium this is (yet). The MAG was generated by researchers at the Hebrew University of Jerusalem from a Red Sea metagenome, and it is of scaffold-level quality.
To download the genome, we will use the command curl:
curl https://ftp.ncbi.nlm.nih.gov/genomes/all/GCA/023/356/555/GCA_023356555.1_ASM2335655v1/GCA_023356555.1_ASM2335655v1_genomic.fna.gz -o GCA_023356555.1_ASM2335655v1_genomic.fna.gz
gunzip GCA_023356555.1_ASM2335655v1_genomic.fna.gz
Now we have one of the most fundamental files that everyone will have to interact with: a FASTA file. We can already check the number of contigs by counting the number of > characters, which appears once per sequence. We can use the command grep for that:
grep -c '>' GCA_023356555.1_ASM2335655v1_genomic.fna
You should see the value 269 in the terminal output. That’s a lot of sequences, or in this case: a lot of contigs. That is already telling us a few things about this genome: it is not a singular contig representing a complete and circular genome. Let’s have a look at the contig headers:
grep '>' GCA_023356555.1_ASM2335655v1_genomic.fna
As you can see, the headers are rather complex, with a lot of information. Unfortunately, these headers are going to be problematic for downstream analysis, particularly because they include non-alphanumeric characters like spaces, dots, pipe (|) and whatnot. And anvi’o knows you would be in trouble if you start working this these headers. Just for fun, you can try to run anvi-gen-contigs-database (we will cover what it does very soon), and you will see an error:
anvi-gen-contigs-database -f GCA_023356555.1_ASM2335655v1_genomic.fna \
-o CONTIGS.db \
-T 2
Config Error: At least one of the deflines in your FASTA File does not comply with the 'simple
deflines' requirement of anvi'o. You can either use the script `anvi-script-
reformat-fasta` to take care of this issue, or read this section in the tutorial
to understand the reason behind this requirement (anvi'o is very upset for
making you do this): http://merenlab.org/2016/06/22/anvio-tutorial-v2/#take-a-
look-at-your-fasta-file
We can use anvi-script-reformat-fasta to simplify the FASTA’s headers with the flag --simplify-names. This command can (optionally) generate a summary report, which is a two column file matching the new and old names of each sequence in the FASTA file. While we are using this command, we can use it to include a specific prefix in the renamed headers with the flag --prefix and filter out short contigs (in this example, smaller than 500 bp) with the flag -l.
anvi-script-reformat-fasta GCA_023356555.1_ASM2335655v1_genomic.fna \
-o Trichodesmium_sp.fa \
--simplify-names \
--prefix Trichodesmium_sp_MAG_R01 \
-r Trichodesmium_sp_reformat_report.txt \
-l 500 \
--seq-type NT
You can use this command to further filter your FASTA file: check the options with the online help page, or by using the flag --help in the terminal.
Generate a contigs database
The contigs-db is a central database in the anvi’o ecosystem. It essentially stores all information about a given set of (nucleotide) sequences from a FASTA file. To make a contigs-db from our reformatted FASTA, you can use the command anvi-gen-contigs-database:
Numerous anvi’o commands can make use of multithreading to speed up computing time. You can either use the flag -T or --num-threads. Check the help page of a command, or use --help to check if multithreading is available.
anvi-gen-contigs-database -f Trichodesmium_sp.fa \
-o Trichodesmium_sp-contigs.db \
-T 2
A few things happen when you generate a contigs-db. First, all DNA sequences are stored in that database. You can retrieve the sequences in FASTA format by using the command anvi-export-contigs. Second, anvi’o uses pyrodigal-gv to identify open-reading frames (also referred to as ‘gene calls’). Pyrodigal-gv is a python implementation of Prodigal with some additional metagenomic models for giant viruses and viruses with alternative genetic codes (see Camargo et al.).
In addition to identifying open-reading frames, anvi’o will predict the amino acid sequence associated with each gene call and store it in that newly made database. If you need to get this information out of the database, you can use the command anvi-export-gene-calls to export the gene calls, and the amino acid sequence for each open-reading frame identified by pyrodigal-gv.
The command anvi-gen-contigs-database also computes the tetra-nucleotide frequency for each contig. To learn more about what it is, check the vocabulary page about tetra-nucleotide frequency.
Annotate single-copy core genes and Ribosomal RNAs
There is a command called anvi-run-hmms, which lets you use Hidden Markov Models (HMMs) to annotate the genes in a contigs-db and store those annotations directly back into the database.
The anvi’o codebase comes with an integrated set of default HMM sources. They include models for 6 Ribosomal RNAs (16S, 23S, 5S, 18S, 28S, and 12S). They also include three sets of single-copy core genes, named Bacteria_71, Archaea_76 and Protista_83. The first two, Bacteria_71 and Archaea_76, are collections of bacterial and archaeal single-copy core genes (SCGs) curated from Mike Lee’s SCG collections first released in GToTree, which is an easy-to-use phylogenomics workflow. Protista_83 is a curated collection of BUSCO SCGs made by Tom Delmont. These sets of HMMs are used in anvi’o to compute the estimated completeness and redundancy of a genome.
To annotate our contigs-db with these HHMs, we can simply run anvi-run-hmms like this:
anvi-run-hmms -c Trichodesmium_sp-contigs.db -T 4
There is an optional flag --also-scan-trnas which uses the program tRNAScan-SE to identify and store information about tRNAs found in your genome. You can also use the command anvi-scan-trnas at any time.
Now is probably a good time to use the command anvi-db-info, which shows you basic information about any anvi’o database:
anvi-db-info Trichodesmium_sp-contigs.db
With this command, you can see which HMMs were already run on that database, but also some basic information like the number of contigs, number of genes called by pyrodigal-gv, and more:
DB Info (no touch)
===============================================
Database Path ................................: Trichodesmium_sp-contigs.db
description ..................................: [Not found, but it's OK]
db_type ......................................: contigs (variant: unknown)
version ......................................: 24
DB Info (no touch also)
===============================================
project_name .................................: Trichodesmium_sp
contigs_db_hash ..............................: hash0d1122fb
split_length .................................: 20000
kmer_size ....................................: 4
num_contigs ..................................: 269
total_length .................................: 6640707
num_splits ...................................: 358
gene_level_taxonomy_source ...................: None
genes_are_called .............................: 1
external_gene_calls ..........................: 0
external_gene_amino_acid_seqs ................: 0
skip_predict_frame ...........................: 0
splits_consider_gene_calls ...................: 1
trna_taxonomy_was_run ........................: 0
trna_taxonomy_database_version ...............: None
reaction_network_ko_annotations_hash .........: None
reaction_network_kegg_database_release .......: None
reaction_network_modelseed_database_sha ......: None
reaction_network_consensus_threshold .........: None
reaction_network_discard_ties ................: None
creation_date ................................: 1760017061.92556
scg_taxonomy_was_run .........................: 1
scg_taxonomy_database_version ................: GTDB: v214.1; Anvi'o: v1
gene_function_sources ........................: COG24_FUNCTION,COG24_PATHWAY,COG24_CATEGORY,KOfam,KEGG_BRITE,KEGG_Class,KEGG_Module
modules_db_hash ..............................: 66e53d49e65a
* Please remember that it is never a good idea to change these values. But in some
cases it may be absolutely necessary to update something here, and a
programmer may ask you to run this program and do it. But even then, you
should be extremely careful.
AVAILABLE GENE CALLERS
===============================================
* 'pyrodigal-gv' (4,820 gene calls)
AVAILABLE FUNCTIONAL ANNOTATION SOURCES
===============================================
* No functional annotations found in this contigs database :/
AVAILABLE HMM SOURCES
===============================================
* 'Archaea_76' (76 models with 34 hits)
* 'Bacteria_71' (71 models with 72 hits)
* 'Protista_83' (83 models with 38 hits)
* 'Ribosomal_RNA_12S' (1 model with 0 hits)
* 'Ribosomal_RNA_16S' (3 models with 0 hits)
* 'Ribosomal_RNA_18S' (1 model with 0 hits)
* 'Ribosomal_RNA_23S' (2 models with 0 hits)
* 'Ribosomal_RNA_28S' (1 model with 0 hits)
* 'Ribosomal_RNA_5S' (5 models with 0 hits)
General summary and metrics
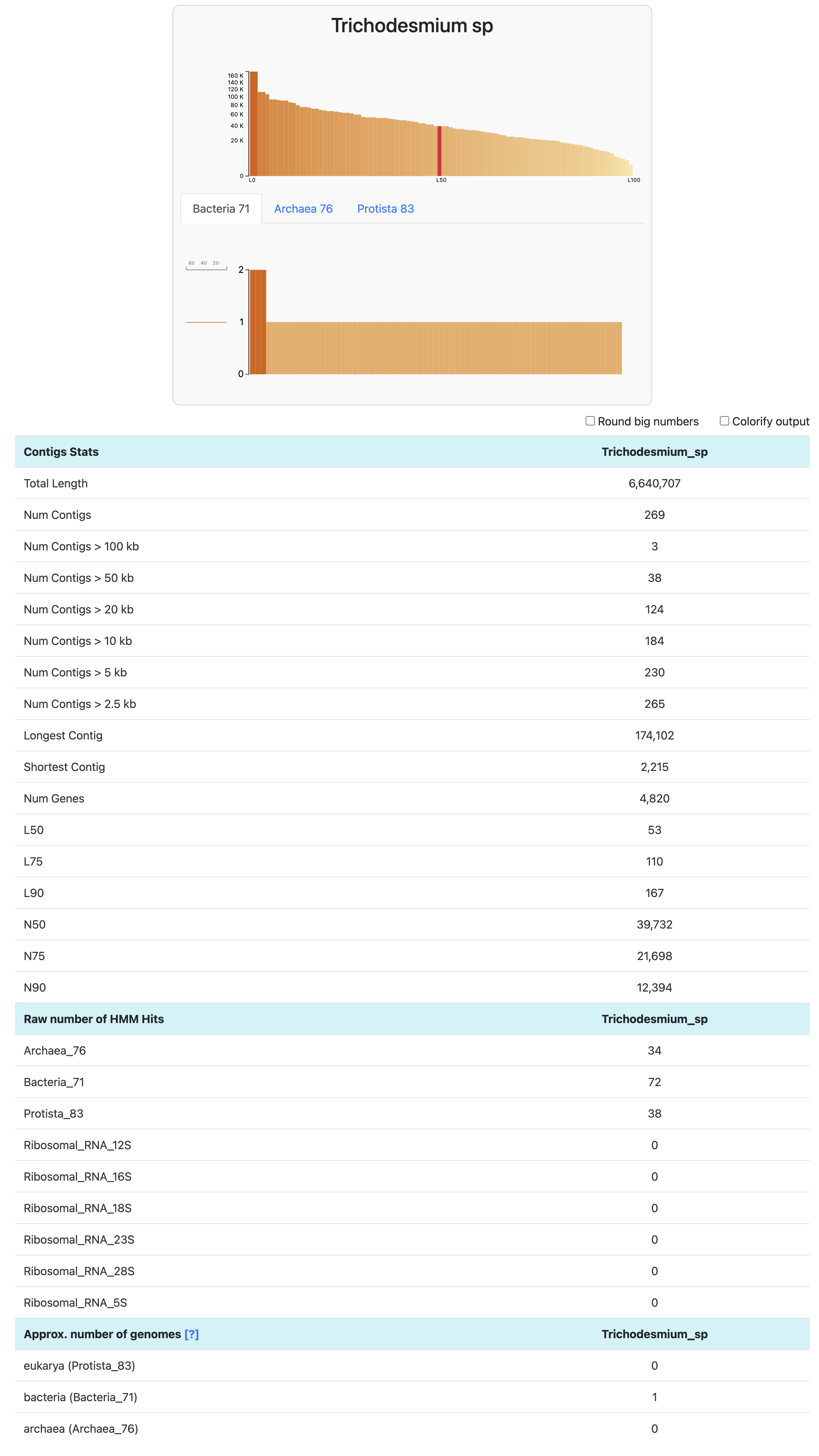
To go one step further than the output generated by anvi-db-info, we can use the command anvi-display-contigs-stats. It allows you to visualize (and export) basic information about one or multiple contigs databases.
anvi-display-contigs-stats Trichodesmium_sp-contigs.db
The approximate number of genomes is an estimate based on the frequency of each single-copy core gene (more info here). It is mostly useful in a metagenomics context, where you expect multiple microbial populations to be included in your set of contigs. This estimate is based on the mode (or most frequently occurring number) of single-copy core genes. Here, we know we generated a contigs-db with a single genome, hence the expected number of genomes = 1 bacteria. GOOD.
You can also export the information displayed on your browser by running anvi-display-contigs-stats with the flag --output-file or -o. Anvi’o will then write a TAB-delimited file with the above values. And if you don’t want the browser page to open at all, you can also use the flag --report-as-text.
anvi-display-contigs-stats Trichodesmium_sp-contigs.db --report-as-text -o Trichodesmium_sp-contigs-stats.txt
Estimate completeness and redundancy
We can use the set of single-copy core genes (SCGs) to estimate the completeness of a genome. The rationale is pretty simple: we expect a set of genes to be systematically present in all genomes only once. So if we find all these genes, we estimate that the genome is complete (100%). We can also report how many SCGs are found in multiple copies, which we refer to as SCG ‘redundancy’.
Why redundancy and not contamination? The presence of multiple copies of SCGs could be indicative of contamination in your genome - i.e., the potential presence of more than one population - but microbial genomes have a bad habit of keeping a few SCGs in multiple copies. So until proven otherwise, anvi’o calls it redundancy and you will decide if you think it is contamination.
We can use the command anvi-estimate-genome-completeness:
anvi-estimate-genome-completeness -c Trichodesmium_sp-contigs.db
bin name |
domain |
confidence |
% completion |
% redundancy |
num_splits |
total length |
|---|---|---|---|---|---|---|
| Trichodesmium_sp-contigs | BACTERIA | 1.0 | 97.18 | 4.23 | 358 | 6640707 |
From the output, we learn that our MAG is estimated to be 97% complete and 4% redundant.
You can also use the flag --output-file or -o to save the output into a text file.
Estimate SCG taxonomy
In this part of the tutorial, we will cover how anvi’o can generate a quick taxonomic estimation for a genome (also works for a metagenome). Of course, in this example, we already know that our genome is from the genus Trichodesmium (because its information on NCBI said so). But we don’t know exactly which species of Trichodesmium it is. You can also imagine doing this step without any prior knowledge about the taxonomy of your microbe.
The way anvi’o estimates taxonomy is by using a subset of genes found in the bacterial and archaeal SCG collections: the ribosomal proteins. These proteins are commonly used to compute phylogenomic trees. Briefly, anvi’o uses the Genome Taxonomy Database (GTDB) and diamond (a fast alternative to NCBI’s BLASTp) to compare your ribosomal proteins to those found in the GTDB collection. Each ribosomal protein thus gets a closest taxonomic match. This is done with the command anvi-run-scg-taxonomy, which stores the taxonomic annotations directly in the contigs-db.
Let’s run anvi-run-scg-taxonomy:
anvi-run-scg-taxonomy -c Trichodesmium_sp-contigs.db -T 4
If this step is not working for you, you may need to run anvi-setup-scg-taxonomy to download and initialize the GTDB SCG database on your machine. You only need to run this command once.
In your terminal, you should see the number of ribosomal proteins that had a match to a homologous protein from the GTDB collection. But you don’t get the actual taxonomy yet. For that, you need to run a second program called anvi-estimate-scg-taxonomy. This command will compute the consensus taxonomic annotation across a set of ribosomal proteins. If we simply provide our contigs-db as the sole input to anvi-estimate-scg-taxonomy, then anvi’o will assume it contains a single genome and compute the consensus across all ribosomal protein matches.
Let’s run anvi-estimate-scg-taxonomy:
anvi-estimate-scg-taxonomy -c Trichodesmium_sp-contigs.db
Contigs DB ...................................: Trichodesmium_sp-contigs.db
Metagenome mode ..............................: False
Estimated taxonomy for "Trichodesmium_sp"
===============================================
+------------------+--------------+-------------------+----------------------------------------------------------------------------------------------------------------------------+
| | total_scgs | supporting_scgs | taxonomy |
+==================+==============+===================+============================================================================================================================+
| Trichodesmium_sp | 22 | 21 | Bacteria / Cyanobacteriota / Cyanobacteriia / Cyanobacteriales / Microcoleaceae / Trichodesmium / Trichodesmium erythraeum |
+------------------+--------------+-------------------+----------------------------------------------------------------------------------------------------------------------------+
Note that the command used a total of 22 SCGs - more specifically, the ribosomal proteins that are part of the SCG collection - and we now have an estimated species name: Trichodesmium erythraeum.
In the output, you can also see supporting_scgs with the number 21. It corresponds to the number of ribosomal proteins which all agreed with the reported consensus taxonomy. It also means that ONE ribosomal protein has a different taxonomic match.
If you are curious, we can run the same command with the flag --debug:
anvi-estimate-scg-taxonomy -c Trichodesmium_sp-contigs.db --debug
Contigs DB ...................................: Trichodesmium_sp-contigs.db
Metagenome mode ..............................: False
* A total of 22 single-copy core genes with taxonomic affiliations were
successfully initialized from the contigs database 🎉 Following shows the
frequency of these SCGs: Ribosomal_L1 (1), Ribosomal_L13 (1), Ribosomal_L14
(1), Ribosomal_L16 (1), Ribosomal_L17 (1), Ribosomal_L19 (1), Ribosomal_L2
(1), Ribosomal_L20 (1), Ribosomal_L21p (1), Ribosomal_L22 (1), Ribosomal_L27A
(1), Ribosomal_L3 (1), Ribosomal_L4 (1), Ribosomal_L5 (1), Ribosomal_S11 (1),
Ribosomal_S15 (1), Ribosomal_S16 (1), Ribosomal_S2 (1), Ribosomal_S6 (1),
Ribosomal_S7 (1), Ribosomal_S8 (1), Ribosomal_S9 (1).
Hits for Trichodesmium_sp
===============================================
+----------------+--------+----------+----------------------------------------------------------------------------------------------------------------------------+
| SCG | gene | pct id | taxonomy |
+================+========+==========+============================================================================================================================+
| Ribosomal_L19 | 2881 | 99.2 | Bacteria / Cyanobacteriota / Cyanobacteriia / Cyanobacteriales / Microcoleaceae / Trichodesmium / Trichodesmium erythraeum |
+----------------+--------+----------+----------------------------------------------------------------------------------------------------------------------------+
| Ribosomal_S2 | 1476 | 99.3 | Bacteria / Cyanobacteriota / Cyanobacteriia / Cyanobacteriales / Microcoleaceae / Trichodesmium / Trichodesmium erythraeum |
+----------------+--------+----------+----------------------------------------------------------------------------------------------------------------------------+
| Ribosomal_L20 | 967 | 99.2 | Bacteria / Cyanobacteriota / Cyanobacteriia / Cyanobacteriales / Microcoleaceae / Trichodesmium / Trichodesmium erythraeum |
+----------------+--------+----------+----------------------------------------------------------------------------------------------------------------------------+
| Ribosomal_S6 | 4430 | 95.7 | Bacteria / Cyanobacteriota / Cyanobacteriia / Cyanobacteriales / Microcoleaceae / Trichodesmium / Trichodesmium erythraeum |
+----------------+--------+----------+----------------------------------------------------------------------------------------------------------------------------+
| Ribosomal_S9 | 1368 | 100.0 | Bacteria / Cyanobacteriota / Cyanobacteriia / Cyanobacteriales / Microcoleaceae / Trichodesmium / Trichodesmium erythraeum |
+----------------+--------+----------+----------------------------------------------------------------------------------------------------------------------------+
| Ribosomal_L13 | 1369 | 100.0 | Bacteria / Cyanobacteriota / Cyanobacteriia / Cyanobacteriales / Microcoleaceae / Trichodesmium / Trichodesmium erythraeum |
+----------------+--------+----------+----------------------------------------------------------------------------------------------------------------------------+
| Ribosomal_L17 | 1371 | 98.3 | Bacteria / Cyanobacteriota / Cyanobacteriia / Cyanobacteriales / Microcoleaceae / Trichodesmium / Trichodesmium erythraeum |
+----------------+--------+----------+----------------------------------------------------------------------------------------------------------------------------+
| Ribosomal_S11 | 1373 | 100.0 | Bacteria / Cyanobacteriota / Cyanobacteriia / Cyanobacteriales / Microcoleaceae / Trichodesmium / Trichodesmium erythraeum |
+----------------+--------+----------+----------------------------------------------------------------------------------------------------------------------------+
| Ribosomal_L21p | 352 | 99.3 | Bacteria / Cyanobacteriota / Cyanobacteriia / Cyanobacteriales / Microcoleaceae / Trichodesmium / Trichodesmium erythraeum |
+----------------+--------+----------+----------------------------------------------------------------------------------------------------------------------------+
| Ribosomal_L27A | 1378 | 100.0 | Bacteria / Cyanobacteriota / Cyanobacteriia / Cyanobacteriales / Microcoleaceae / Trichodesmium / Trichodesmium erythraeum |
+----------------+--------+----------+----------------------------------------------------------------------------------------------------------------------------+
| Ribosomal_S8 | 1382 | 100.0 | Bacteria / Cyanobacteriota / Cyanobacteriia / Cyanobacteriales / Microcoleaceae / Trichodesmium / Trichodesmium erythraeum |
+----------------+--------+----------+----------------------------------------------------------------------------------------------------------------------------+
| Ribosomal_L5 | 1383 | 99.4 | Bacteria / Cyanobacteriota / Cyanobacteriia / Cyanobacteriales / Microcoleaceae / Trichodesmium / Trichodesmium erythraeum |
+----------------+--------+----------+----------------------------------------------------------------------------------------------------------------------------+
| Ribosomal_L14 | 1385 | 100.0 | Bacteria / Cyanobacteriota / Cyanobacteriia / Cyanobacteriales / Microcoleaceae / Trichodesmium / |
+----------------+--------+----------+----------------------------------------------------------------------------------------------------------------------------+
| Ribosomal_L16 | 1388 | 100.0 | Bacteria / Cyanobacteriota / Cyanobacteriia / Cyanobacteriales / Microcoleaceae / Trichodesmium / Trichodesmium erythraeum |
+----------------+--------+----------+----------------------------------------------------------------------------------------------------------------------------+
| Ribosomal_L22 | 1390 | 100.0 | Bacteria / Cyanobacteriota / Cyanobacteriia / Cyanobacteriales / Microcoleaceae / Trichodesmium / Trichodesmium erythraeum |
+----------------+--------+----------+----------------------------------------------------------------------------------------------------------------------------+
| Ribosomal_S15 | 1646 | 100.0 | Bacteria / Cyanobacteriota / Cyanobacteriia / Cyanobacteriales / Microcoleaceae / Trichodesmium / Trichodesmium erythraeum |
+----------------+--------+----------+----------------------------------------------------------------------------------------------------------------------------+
| Ribosomal_L2 | 1392 | 97.9 | Bacteria / Cyanobacteriota / Cyanobacteriia / Cyanobacteriales / Microcoleaceae / Trichodesmium / Trichodesmium erythraeum |
+----------------+--------+----------+----------------------------------------------------------------------------------------------------------------------------+
| Ribosomal_L4 | 1394 | 99.5 | Bacteria / Cyanobacteriota / Cyanobacteriia / Cyanobacteriales / Microcoleaceae / Trichodesmium / Trichodesmium erythraeum |
+----------------+--------+----------+----------------------------------------------------------------------------------------------------------------------------+
| Ribosomal_L3 | 1395 | 98.2 | Bacteria / Cyanobacteriota / Cyanobacteriia / Cyanobacteriales / Microcoleaceae / Trichodesmium / Trichodesmium erythraeum |
+----------------+--------+----------+----------------------------------------------------------------------------------------------------------------------------+
| Ribosomal_S7 | 3771 | 100.0 | Bacteria / Cyanobacteriota / Cyanobacteriia / Cyanobacteriales / Microcoleaceae / Trichodesmium / Trichodesmium erythraeum |
+----------------+--------+----------+----------------------------------------------------------------------------------------------------------------------------+
| Ribosomal_L1 | 2877 | 100.0 | Bacteria / Cyanobacteriota / Cyanobacteriia / Cyanobacteriales / Microcoleaceae / Trichodesmium / Trichodesmium erythraeum |
+----------------+--------+----------+----------------------------------------------------------------------------------------------------------------------------+
| Ribosomal_S16 | 1407 | 100.0 | Bacteria / Cyanobacteriota / Cyanobacteriia / Cyanobacteriales / Microcoleaceae / Trichodesmium / Trichodesmium erythraeum |
+----------------+--------+----------+----------------------------------------------------------------------------------------------------------------------------+
| CONSENSUS | -- | -- | Bacteria / Cyanobacteriota / Cyanobacteriia / Cyanobacteriales / Microcoleaceae / Trichodesmium / Trichodesmium erythraeum |
+----------------+--------+----------+----------------------------------------------------------------------------------------------------------------------------+
Estimated taxonomy for "Trichodesmium_sp"
===============================================
+------------------+--------------+-------------------+----------------------------------------------------------------------------------------------------------------------------+
| | total_scgs | supporting_scgs | taxonomy |
+==================+==============+===================+============================================================================================================================+
| Trichodesmium_sp | 22 | 21 | Bacteria / Cyanobacteriota / Cyanobacteriia / Cyanobacteriales / Microcoleaceae / Trichodesmium / Trichodesmium erythraeum |
+------------------+--------------+-------------------+----------------------------------------------------------------------------------------------------------------------------+
In this more comprehensive output, you can see the detail of each ribosomal protein’s closest match to the GTDB SCG database: which protein, the percent identity to an homologous gene in the GTDB collection, and the associated taxonomy. You can see that ribosomal protein L14 has a taxonomy that is not resolved at the species level, but only at the genus level. So, the only ribosomal protein that did not agree with the final taxonomy was actually matching the same genus. I would not be worried at all about potential contamination, at least at this stage.
The way anvi’o estimates taxonomy is not perfect. In fact, it is limited to Bacteria and Archaea (genomes found in GTDB). But it is fast and relatively accurate to give you an idea of the taxonomy of one or more genomes.
If you are working with a metagenome, i.e. with more than one population, anvi’o can either compute the taxonomy per bin (if available), or without binning! In the second case, anvi’o will pick the most commonly found ribosomal protein and report their closest taxonomic matches. This is a great way to quickly get an idea of the taxonomic composition of a metagenome prior to binning.
Functional annotations
There are many databases that one can use to assign function to a set of genes. You may be familiar with NCBI’s COG (Clusters of Orthologous Genes) database, the KEGG (Kyoto Encyclopedia of Genes and Genomes) database of KEGG Ortholog families (KOfam), or Pfam (the Protein Family database), which can be used for annotating general functions. Other databases out there are more specific, like CAZymes which focuses on enzymes associated with the synthesis, metabolism and recognition of carbohydrate, or PHROGs which focuses on viral functions.
Anvi’o has a few commands that allow you to annotate the open-reading frames in your contigs-db with several different databases. Each of them comes with a setup command that you need to run once to download the appropriate database on your machine:
| Database | Setup command | Run command |
|---|---|---|
| NCBI COGs | anvi-setup-ncbi-cogs | anvi-run-ncbi-cogs |
| KEGG KOfam | anvi-setup-kegg-data | anvi-run-kegg-kofams |
| Pfam | anvi-setup-pfams | anvi-run-pfams |
| CAZymes | anvi-setup-cazymes | anvi-run-cazymes |
If your favorite annotation database is not represented here, you have a few options:
- Short-term solution: run the annotation outside of anvi’o. You can export the gene calls with anvi-export-gene-calls, run your annotation with a third party software, then import the annotations back into your contigs-db with anvi-import-functions.
- Community-level solution: find an anvi’o developer and tell them about your passion for XXX functional database and hope they make a new command called
anvi-run-XXX. We seriously encourage you to use the anvi’o github page to write an issue describing the need for a new functional annotation database in anvi’o. Then, anyone with time and skill can try to implement it. If you find an existing issue discussing something you want in anvi’o, please raise your voice, write a comment, and let the developers know that you would like to see a feature in anvi’o. - Developer-level solution: write a new program to annotate a contigs-db with your favorite database, and submit a pull request on the anvi’o github page.
For now, we will use the COG, KEGG, and Pfam databases on our Trichodesmium genome. anvi-run-ncbi-cogs will be relatively fast (it uses diamond to find homologous hits in the database), while anvi-run-kegg-kofams will take much longer as the KOfam database is quite large (this program uses HMMs and HMMER in the background). The Pfam database also uses HMMs, but the domain-level models are a bit smaller so anvi-run-pfams is relatively quick to run.
If you know your machine can use more threads, feel free to change the flag -T 4 to another number.
# will take ~1 min
anvi-run-ncbi-cogs -c Trichodesmium_sp-contigs.db -T 4
# will take ~8 min
anvi-run-kegg-kofams -c Trichodesmium_sp-contigs.db -T 4
# will take ~1 min
anvi-run-pfams -c Trichodesmium_sp-contigs.db -T 4
As you can see in the terminal output from the commands above, there are no output files. All the annotations are stored into the contigs-db. You can use anvi-db-info to check which functional annotations are already in an existing contigs-db (including manually imported ones).
# check which annotations were run on our contigs database
anvi-db-info Trichodesmium_sp-contigs.db
DB Info (no touch)
===============================================
Database Path ................................: Trichodesmium_sp-contigs.db
description ..................................: [Not found, but it's OK]
db_type ......................................: contigs (variant: unknown)
version ......................................: 24
DB Info (no touch also)
===============================================
project_name .................................: Trichodesmium_sp
contigs_db_hash ..............................: hash98a3e869
split_length .................................: 20000
kmer_size ....................................: 4
num_contigs ..................................: 269
total_length .................................: 6640707
num_splits ...................................: 358
gene_level_taxonomy_source ...................: None
genes_are_called .............................: 1
external_gene_calls ..........................: 0
external_gene_amino_acid_seqs ................: 0
skip_predict_frame ...........................: 0
splits_consider_gene_calls ...................: 1
trna_taxonomy_was_run ........................: 0
trna_taxonomy_database_version ...............: None
reaction_network_ko_annotations_hash .........: None
reaction_network_kegg_database_release .......: None
reaction_network_modelseed_database_sha ......: None
reaction_network_consensus_threshold .........: None
reaction_network_discard_ties ................: None
creation_date ................................: 1759736966.76845
scg_taxonomy_was_run .........................: 1
scg_taxonomy_database_version ................: GTDB: v214.1; Anvi'o: v1
gene_function_sources ........................: COG24_CATEGORY,KEGG_BRITE,KEGG_Module,COG24_PATHWAY,KOfam,COG24_FUNCTION,KEGG_Class
modules_db_hash ..............................: a2b5bde358bb
* Please remember that it is never a good idea to change these values. But in some
cases it may be absolutely necessary to update something here, and a
programmer may ask you to run this program and do it. But even then, you
should be extremely careful.
AVAILABLE GENE CALLERS
===============================================
* 'pyrodigal-gv' (4,820 gene calls)
AVAILABLE FUNCTIONAL ANNOTATION SOURCES
===============================================
* COG24_CATEGORY (3,098 annotations)
* COG24_FUNCTION (3,098 annotations)
* COG24_PATHWAY (858 annotations)
* KEGG_BRITE (1,939 annotations)
* KEGG_Class (449 annotations)
* KEGG_Module (449 annotations)
* KOfam (1,941 annotations)
AVAILABLE HMM SOURCES
===============================================
* 'Archaea_76' (76 models with 34 hits)
* 'Bacteria_71' (71 models with 72 hits)
* 'Protista_83' (83 models with 38 hits)
* 'Ribosomal_RNA_12S' (1 model with 0 hits)
* 'Ribosomal_RNA_16S' (3 models with 0 hits)
* 'Ribosomal_RNA_18S' (1 model with 0 hits)
* 'Ribosomal_RNA_23S' (2 models with 0 hits)
* 'Ribosomal_RNA_28S' (1 model with 0 hits)
* 'Ribosomal_RNA_5S' (5 models with 0 hits)
You can also use anvi-export-functions to get a tab-delimited file of all the annotations from a given annotation source (or multiple).
# export functions for KOfam and COG24_FUCTION
anvi-export-functions -c Trichodesmium_sp-contigs.db --annotation-sources KOfam,COG24_FUNCTION -o functional_annotations.txt
The output table from anvi-export-functions looks like this:
gene_callers_id |
source |
accession |
function |
e_value |
|---|---|---|---|---|
| 0 | COG24_FUNCTION | COG3293 | Transposase | 5.51e-12 |
| 2 | COG24_FUNCTION | COG4451 | Ribulose bisphosphate carboxylase small subunit (RbcS) (PDB:2YBV) | 2.02e-63 |
| 4 | COG24_FUNCTION | COG1850 | Ribulose 1,5-bisphosphate carboxylase, large subunit, or a RuBisCO-like protein (RbcL) (PDB:2YBV) | 0 |
| .. | .. | .. | .. | .. |
| 4765 | KOfam | K02030 | polar amino acid transport system substrate-binding protein | 1.7e-26 |
| 4769 | KOfam | K07494 | putative transposase | 3.6e-16 |
| 4813 | KOfam | K11524 | positive phototaxis protein PixI | 4e-37 |
You can search for your favorite function. As we discussed above, Trichodesmium is known for its ability to fix nitrogen, so you can look for the NifH gene, which is a marker gene for nitrogen fixation. Here is how to do that with a simple grep command:
grep NifH functional_annotations.txt
3709 COG24_FUNCTION COG1348 Nitrogenase ATPase subunit NifH/coenzyme F430 biosynthesis subunit CfbC (NifH/CfbC) (PDB:1CP2) (PUBMED:28225763) 1.5e-197
4020 COG24_FUNCTION COG1348 Nitrogenase ATPase subunit NifH/coenzyme F430 biosynthesis subunit CfbC (NifH/CfbC) (PDB:1CP2) (PUBMED:28225763) 4.81e-167
4020 KOfam K02588 nitrogenase iron protein NifH 2.9e-144
This is unexpected: there are two genes (caller id: 3709 and 4020) with the NifH annotation. On of them has both an annotation by COG and KEGG; that’s good, we like consistency. But we expect to find only one copy of the NifH gene, and the other gene only has a COG annotation. Turns out that the COG annotation is not very reliable and in the paper, Tom noticed that the COG was wrongly annotating a Ferredoxin as NifH:
For instance, we found that genes with COG20 function incorrectly annotated as “Nitrogenase ATPase subunit NifH/coenzyme F430 biosynthesis subunit CfbC” correspond, in reality, to “ferredoxin: protochlorophyllide reductase.”
An alternative search strategy: you can use anvi-search-functions, this time using only the KOfam annotation source:
anvi-search-functions -c Trichodesmium_sp-contigs.db \
--search-term NifH \
--annotation-sources KOfam \
--output-file NifH_search.txt \
--full-report NifH_full_report.txt
The first output file called NifH_search.txt only contains the name of the contigs where a gene with a matching search term was found. And the second file, NifH_full_report.txt, is more comprehensive:
cat NifH_full_report.txt
gene_callers_id source accession function search_term contigs
4020 KOfam K02588 nitrogenase iron protein NifH NifH Trichodesmium_sp_MAG_R01_000000000230_split_00006
We’ve found our marker gene for nitrogen fixation, which is a good sign given that this MAG seems to be a T. erythraeum genome, and T. erythraeum is known to fix nitrogen.
Working with multiple genomes
Now that we know how to do basic genomic analysis using a single genome, we can try to do the same using a few more genomes. In the directory 00_FASTA_GENOMES, you will find seven FASTA files containing the reference genomes and MAGs from Tom’s paper:
$ ls 00_DATA/fasta
MAG_Candidatus_Trichodesmium_miru.fa MAG_Trichodesmium_erythraeum.fa MAG_Trichodesmium_thiebautii_Indian.fa Trichodesmium_thiebautii_H9_4.fa
MAG_Candidatus_Trichodesmium_nobis.fa MAG_Trichodesmium_thiebautii_Atlantic.fa Trichodesmium_erythraeum_IMS101.fa
We will create as many contigs databases as we have genomes in this directory. We will then annotate them in a similar fashion as when working with a single genome.
To avoid too much manual labor, we’ll use BASH loops to automate the process. The loops will be a bit easier to write (and understand) if we have a text file of genome names to iterate over. So first, let’s create a simple text file that contains the names of our genomes. The following BASH command will list the content of the 00_FASTA_GENOMES directory and will only keep the part of the file name before the .fa extension, a.k.a. only the name of each genome:
ls 00_DATA/fasta | cut -d "." -f1 > genomes.txt
The second thing to do is to make sure our FASTA files are properly formatted. Fortunately for you, we provided genomes with anvi’o compatible headers. If you don’t believe me (and you should never believe me, and always check your data), then have a look at them.
The next step is to generate contigs-db for each of our genomes with the following BASH loop:
while read genome
do
anvi-gen-contigs-database -f 00_DATA/fasta/${genome}.fa \
-o ${genome}-contigs.db \
-T 4
done < genomes.txt
Now we can annotate these genomes with anvi-run-hmms, anvi-run-ncbi-cogs, anvi-run-kegg-kofams, and anvi-run-pfams. Some of those annotation commands can take a while, so if you don’t want to wait, click the Show/Hide box below to instead get the already-annotated contigs databases from the datapack:
Show/Hide Option 1: Copy pre-annotated databases
This command overwrite the databases in your current working directory with our pre-annotated ones from the 00_DATA folder:
cp 00_DATA/contigs/*-contigs.db .
But if you do want to run the annotations yourself, I encourage you to try writing the loop before checking the answer in the following Show/Hide box:
Show/Hide Option 2: Loop to annotate all databases
while read genome
do
echo "working on $genome"
anvi-run-hmms -c ${genome}-contigs.db -T 4
anvi-run-ncbi-cogs -c ${genome}-contigs.db -T 4
anvi-run-kegg-kofams -c ${genome}-contigs.db -T 4
anvi-run-pfams -c ${genome}-contigs.db -T 4
anvi-run-scg-taxonomy -c ${genome}-contigs.db -T 4
done < genomes.txt
You don’t always need to write a loop to process multiple contigs databases. Some commands, like anvi-estimate-genome-completeness, can work on multiple input databases and will create individual outputs for each one. In these cases, you can use a special table that we call an ‘external genomes’ file. It is a simple two-column table containing the name of each contigs-db and the path to each database.
You can make that table yourself easily, but if you are like me - too lazy to do that by yourself - then you can use anvi-script-gen-genomes-file:
anvi-script-gen-genomes-file --input-dir . -o external-genomes.txt
And here is how this external-genomes file looks like:
name |
contigs_db_path |
|---|---|
| MAG_Candidatus_Trichodesmium_miru | /path/to/MAG_Candidatus_Trichodesmium_miru-contigs.db |
| MAG_Candidatus_Trichodesmium_nobis | /path/to/MAG_Candidatus_Trichodesmium_nobis-contigs.db |
| MAG_Trichodesmium_erythraeum | /path/to/MAG_Trichodesmium_erythraeum-contigs.db |
| MAG_Trichodesmium_thiebautii_Atlantic | /path/to/MAG_Trichodesmium_thiebautii_Atlantic-contigs.db |
| MAG_Trichodesmium_thiebautii_Indian | /path/to/MAG_Trichodesmium_thiebautii_Indian-contigs.db |
| Trichodesmium_erythraeum_IMS101 | /path/to/Trichodesmium_erythraeum_IMS101-contigs.db |
| Trichodesmium_sp | /path/to/Trichodesmium_sp-contigs.db |
| Trichodesmium_thiebautii_H9_4 | /path/to/Trichodesmium_thiebautii_H9_4-contigs.db |
Now we can use this file as the input for commands like anvi-estimate-genome-completeness and anvi-estimate-scg-taxonomy:
anvi-estimate-genome-completeness -e external-genomes.txt
+---------------------------------------+----------+--------------+----------------+----------------+--------------+----------------+
| genome name | domain | confidence | % completion | % redundancy | num_splits | total length |
+=======================================+==========+==============+================+================+==============+================+
| MAG_Candidatus_Trichodesmium_miru | BACTERIA | 1 | 98.59 | 7.04 | 665 | 5425804 |
+---------------------------------------+----------+--------------+----------------+----------------+--------------+----------------+
| MAG_Candidatus_Trichodesmium_nobis | BACTERIA | 0.9 | 95.77 | 2.82 | 768 | 6101640 |
+---------------------------------------+----------+--------------+----------------+----------------+--------------+----------------+
| MAG_Trichodesmium_erythraeum | BACTERIA | 1 | 97.18 | 5.63 | 644 | 6773488 |
+---------------------------------------+----------+--------------+----------------+----------------+--------------+----------------+
| MAG_Trichodesmium_thiebautii_Atlantic | BACTERIA | 0.9 | 85.92 | 7.04 | 1136 | 5948726 |
+---------------------------------------+----------+--------------+----------------+----------------+--------------+----------------+
| MAG_Trichodesmium_thiebautii_Indian | BACTERIA | 1 | 94.37 | 2.82 | 1394 | 6834732 |
+---------------------------------------+----------+--------------+----------------+----------------+--------------+----------------+
| Trichodesmium_erythraeum_IMS101 | BACTERIA | 1 | 97.18 | 7.04 | 386 | 7750108 |
+---------------------------------------+----------+--------------+----------------+----------------+--------------+----------------+
| Trichodesmium_sp | BACTERIA | 1 | 97.18 | 4.23 | 358 | 6640707 |
+---------------------------------------+----------+--------------+----------------+----------------+--------------+----------------+
| Trichodesmium_thiebautii_H9_4 | BACTERIA | 0.8 | 71.83 | 12.68 | 201 | 3286556 |
+---------------------------------------+----------+--------------+----------------+----------------+--------------+----------------+
Note that Trichodesmium_thiebautii_H9_4 appears to have quite a low completion estimate and also a rather large redundancy estimate. Just from these two values, we could say that the quality of that MAG is lower than all other genomes in our collection. You may also notice that it has a much smaller genome size (last column).
Let’s try estimating the taxonomy of all our genomes at once with anvi-estimate-scg-taxonomy:
anvi-estimate-scg-taxonomy -e external-genomes.txt -o taxonomy_multi_genomes.txt
Num genomes ..................................: 8
Taxonomic level of interest ..................: (None specified by the user, so 'all levels')
Output file path .............................: taxonomy_multi_genomes.txt
Output raw data ..............................: False
SCG coverages will be computed? ..............: False
* Your (meta)genome file DOES NOT contain profile databases, and you haven't asked
anvi'o to work in `--metagenome-mode`. Your contigs databases will be treated
as genomes rather than metagenomes.
Output file ..................................: taxonomy_multi_genomes.txt
And here is the output:
name |
total_scgs |
supporting_scgs |
t_domain |
t_phylum |
t_class |
t_order |
t_family |
t_genus |
t_species |
|---|---|---|---|---|---|---|---|---|---|
| MAG_Candidatus_Trichodesmium_miru | 22 | 15 | Bacteria | Cyanobacteriota | Cyanobacteriia | Cyanobacteriales | Microcoleaceae | Trichodesmium | Trichodesmium sp023356515 |
| MAG_Candidatus_Trichodesmium_nobis | 20 | 13 | Bacteria | Cyanobacteriota | Cyanobacteriia | Cyanobacteriales | Microcoleaceae | Trichodesmium | Trichodesmium sp023356535 |
| MAG_Trichodesmium_erythraeum | 22 | 21 | Bacteria | Cyanobacteriota | Cyanobacteriia | Cyanobacteriales | Microcoleaceae | Trichodesmium | Trichodesmium erythraeum |
| MAG_Trichodesmium_thiebautii_Atlantic | 21 | 21 | Bacteria | Cyanobacteriota | Cyanobacteriia | Cyanobacteriales | Microcoleaceae | Trichodesmium | Trichodesmium sp023356605 |
| MAG_Trichodesmium_thiebautii_Indian | 22 | 21 | Bacteria | Cyanobacteriota | Cyanobacteriia | Cyanobacteriales | Microcoleaceae | Trichodesmium | Trichodesmium sp023356605 |
| Trichodesmium_erythraeum_IMS101 | 22 | 21 | Bacteria | Cyanobacteriota | Cyanobacteriia | Cyanobacteriales | Microcoleaceae | Trichodesmium | Trichodesmium erythraeum |
| Trichodesmium_sp | 22 | 21 | Bacteria | Cyanobacteriota | Cyanobacteriia | Cyanobacteriales | Microcoleaceae | Trichodesmium | Trichodesmium erythraeum |
| Trichodesmium_thiebautii_H9_4 | 19 | 18 | Bacteria | Cyanobacteriota | Cyanobacteriia | Cyanobacteriales | Microcoleaceae | Trichodesmium | Trichodesmium sp023356605 |
You can see that a lot of the MAGs match to unnamed species in GTDB – even though we already know what most of them are, those names haven’t propagated to the GTDB database yet. This especially makes sense for the candidate species T. miru and T. nobis. Regardless, all the T. thiebautii genomes have the same closest match to T. sp023356535.
A note on anvi’o workflows There are a few built-in snakemake workflows in anvi’o that can be used with the program anvi-run-workflow. We regularly use these workflows for routine analyses, like generating contigs databases and running functional annotations. That is exactly the purpose of the ‘contigs’ workflow.
You don’t need to know anything about snakemake to use these workflows. For instance, for the ‘contigs’ workflow, all you need is two input files:
- A fasta-txt file, which is basically a two-column table with the name and path to each FASTA file that you want to turn into a contigs-db
- A workflow-config file (which you can get from anvi-run-workflow) in which you can specify which commands you want to run and parameters for each command.
This automation sounds like a nice plug-and-play analysis pipeline - and it is - but it requires you to know exactly what you want to run. You are still the chef.
Want to learn more? Check out this other tutorial on running workflows in anvi’o.
Working with one (or more) metagenomes
If you are working with one, or more, metagenomic assemblies, you can use and run the same commands that we ran on individual genomes. This includes anvi-gen-contigs-database, anvi-run-hmms, anvi-run-scg-taxonomy, anvi-run-kegg-kofams, etc.
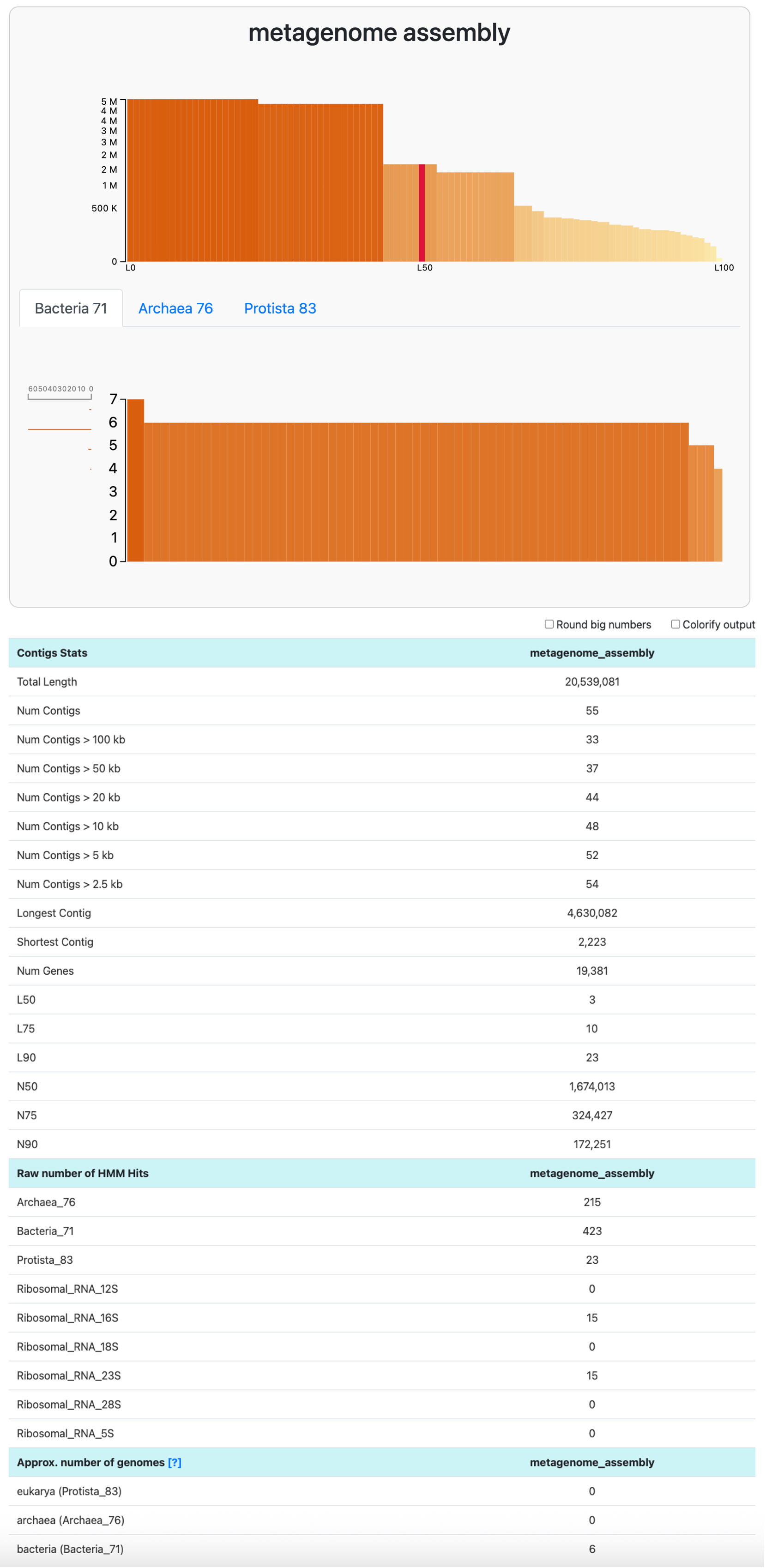
In the datapack, you will find the contigs-db of a mock metagenome assembly that we made for you. With anvi-display-contigs-stats, we can learn about the count and length of contigs in the assembly, as well as the number of expected genomes:
anvi-display-contigs-stats 00_DATA/metagenome/sample01-contigs.db
This assembly is estimated to contain six populations. To learn more about the composition of this metagenome, we can use anvi-estimate-scg-taxonomy with the flag --metagenome-mode. In this mode, anvi’o will not try to compute the consensus taxonomy of every ribosomal protein as it does by default. Instead, it will report the taxonomy of all the genes matching to the most abundant ribosomal protein:
anvi-estimate-scg-taxonomy -c 00_DATA/metagenome/sample01-contigs.db --metagenome-mode
Contigs DB ...................................: 00_DATA/sample01-contigs.db
Metagenome mode ..............................: True
SCG for metagenome ...........................: None
* A total of 132 single-copy core genes with taxonomic affiliations were
successfully initialized from the contigs database 🎉 Following shows the
frequency of these SCGs: Ribosomal_L1 (6), Ribosomal_L13 (6), Ribosomal_L14
(6), Ribosomal_L16 (6), Ribosomal_L17 (6), Ribosomal_L19 (6), Ribosomal_L2
(6), Ribosomal_L20 (6), Ribosomal_L21p (6), Ribosomal_L22 (6), Ribosomal_L27A
(6), Ribosomal_L3 (6), Ribosomal_L4 (6), Ribosomal_L5 (6), Ribosomal_S11 (6),
Ribosomal_S15 (6), Ribosomal_S16 (6), Ribosomal_S2 (6), Ribosomal_S6 (6),
Ribosomal_S7 (6), Ribosomal_S8 (6), Ribosomal_S9 (6).
WARNING
===============================================
Anvi'o automatically set 'Ribosomal_L1' to be THE single-copy core gene to
survey your metagenome for its taxonomic composition. If you are not happy with
that, you could change it with the parameter `--scg-name-for-metagenome-mode`.
Taxa in metagenome "metagenome_assembly"
===============================================
+----------------------------------------+--------------------+--------------------------------------------------------------------------------------------------------------------------+
| | percent_identity | taxonomy |
+========================================+====================+==========================================================================================================================+
| metagenome_assembly_Ribosomal_L1_10579 | 100 | Bacteria / Pseudomonadota / Gammaproteobacteria / Pseudomonadales / Oleiphilaceae / Marinobacter / Marinobacter salarius |
+----------------------------------------+--------------------+--------------------------------------------------------------------------------------------------------------------------+
| metagenome_assembly_Ribosomal_L1_12451 | 100 | Bacteria / Cyanobacteriota / Cyanobacteriia / PCC-6307 / Cyanobiaceae / Prochlorococcus_A / |
+----------------------------------------+--------------------+--------------------------------------------------------------------------------------------------------------------------+
| metagenome_assembly_Ribosomal_L1_15984 | 100 | Bacteria / Pseudomonadota / Alphaproteobacteria / Rhizobiales / Stappiaceae / Roseibium / Roseibium aggregatum |
+----------------------------------------+--------------------+--------------------------------------------------------------------------------------------------------------------------+
| metagenome_assembly_Ribosomal_L1_3513 | 99.6 | Bacteria / Pseudomonadota / Gammaproteobacteria / Enterobacterales_A / Alteromonadaceae / Alteromonas / |
+----------------------------------------+--------------------+--------------------------------------------------------------------------------------------------------------------------+
| metagenome_assembly_Ribosomal_L1_5126 | 100 | Bacteria / Pseudomonadota / Alphaproteobacteria / HIMB59 / HIMB59 / HIMB59 / HIMB59 sp000299115 |
+----------------------------------------+--------------------+--------------------------------------------------------------------------------------------------------------------------+
| metagenome_assembly_Ribosomal_L1_5655 | 99.6 | Bacteria / Pseudomonadota / Alphaproteobacteria / Pelagibacterales / Pelagibacteraceae / Pelagibacter / |
+----------------------------------------+--------------------+--------------------------------------------------------------------------------------------------------------------------+
Alternatively, you can tell anvi’o you want to use another ribosomal protein. The output of the above comment is already telling us that each ribosomal protein is present six times, so any gene should do the job. You can also use the flag --report-scg-frequencies, which will write these frequencies into a text file.
A matrix output with multiple metagenomes
If you have multiple metagenomes, you can use the flag --matrix to get an output file that looks like this (example output at the genus level):
| taxon | sample01 | sample02 | sample03 | sample04 | sample05 | sample06 |
|---|---|---|---|---|---|---|
| Prochlorococcus | 1.00 | 1.00 | 1.00 | 1.00 | 1.00 | 1.00 |
| Synechococcus | 1.00 | 0 | 1.00 | 1.00 | 0 | 1.00 |
| Pelagibacter | 1.00 | 1.00 | 1.00 | 1.00 | 1.00 | 1.00 |
| SAR86 | 1.00 | 1.00 | 0 | 1.00 | 1.00 | 1.00 |
| SAR92 | 0 | 1.00 | 1.00 | 0 | 1.00 | 1.00 |
| SAR116 | 1.00 | 0 | 1.00 | 1.00 | 0 | 1.00 |
| Roseobacter | 0 | 1.00 | 1.00 | 0 | 1.00 | 0 |
The next chapter
If you want to immediately move on to the next chapter of this tutorial, here is the link:
If you want to go back to the main page of the tutorial instead, click here.
If you have any questions about this exercise, or have ideas to make it better, please feel free to get in touch with the anvi’o community through our Discord server: